Explain the formation of ATP in electron transport chain with diagram, mention ATP synthesis sites and inhibitors of ETC uncoupled of etc components of respiratory chain
Reading File
Finding Sources
Finding Sources
Searching Images
electron transport chain ATP synthesis mitochondria diagram
Reading File
Reading File
Finding Sources
Loading Image
Reading File
Finding Sources
Finding Sources
Reading File
Reading File
Finding Sources
Reading File
Loading Image
Loading Image
Loading Image
Reading File
I have all the content needed for a comprehensive answer. Let me now compose the full response:
Electron Transport Chain (ETC) & ATP Synthesis
Overview
Energy-rich molecules (glucose, fatty acids, amino acids) are oxidized through metabolic pathways (glycolysis, β-oxidation, TCA cycle) to generate NADH and FADH₂. These reduced coenzymes donate electrons to the ETC, releasing energy that is captured as ATP. This coupling of electron transport to ATP synthesis is called oxidative phosphorylation (OXPHOS).
Diagram 1 — Components of the Respiratory Chain
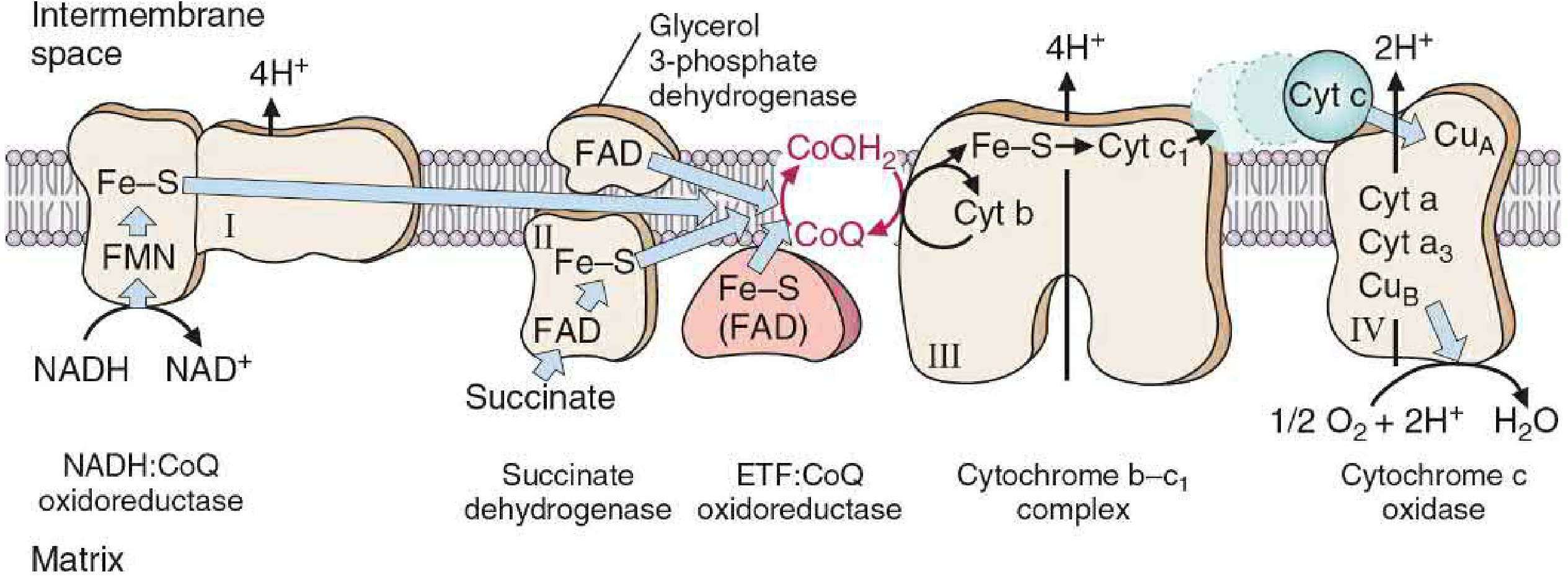
Components of the Respiratory Chain (Complexes I–IV)
| Complex | Name | Prosthetic Groups | Electrons From | Protons Pumped |
|---|---|---|---|---|
| I | NADH:CoQ oxidoreductase (NADH dehydrogenase) | FMN, Fe-S centers | NADH → CoQ | 4 H⁺ |
| II | Succinate:CoQ oxidoreductase (Succinate dehydrogenase) | FAD, Fe-S centers | FADH₂ → CoQ | 0 H⁺ |
| III | CoQH₂:Cytochrome c oxidoreductase (Cytochrome bc₁) | Cyt b, Fe-S, Cyt c₁ | CoQ → Cyt c | 4 H⁺ |
| IV | Cytochrome c oxidase | Cyt a, Cyt a₃, CuA, CuB | Cyt c → O₂ → H₂O | 2 H⁺ |
| V | ATP synthase (F₁F₀-ATPase) | — | H⁺ gradient → ATP | — |
Mobile carriers (not fixed in complexes):
- Coenzyme Q (CoQ / Ubiquinone): Lipid-soluble; shuttles electrons from Complex I and II (and glycerol-3-phosphate dehydrogenase, acyl-CoA dehydrogenase) to Complex III
- Cytochrome c: Water-soluble; peripheral to the inner mitochondrial membrane; shuttles electrons from Complex III to Complex IV
Diagram 2 — ETC with Proton Pumping & ATP Synthesis
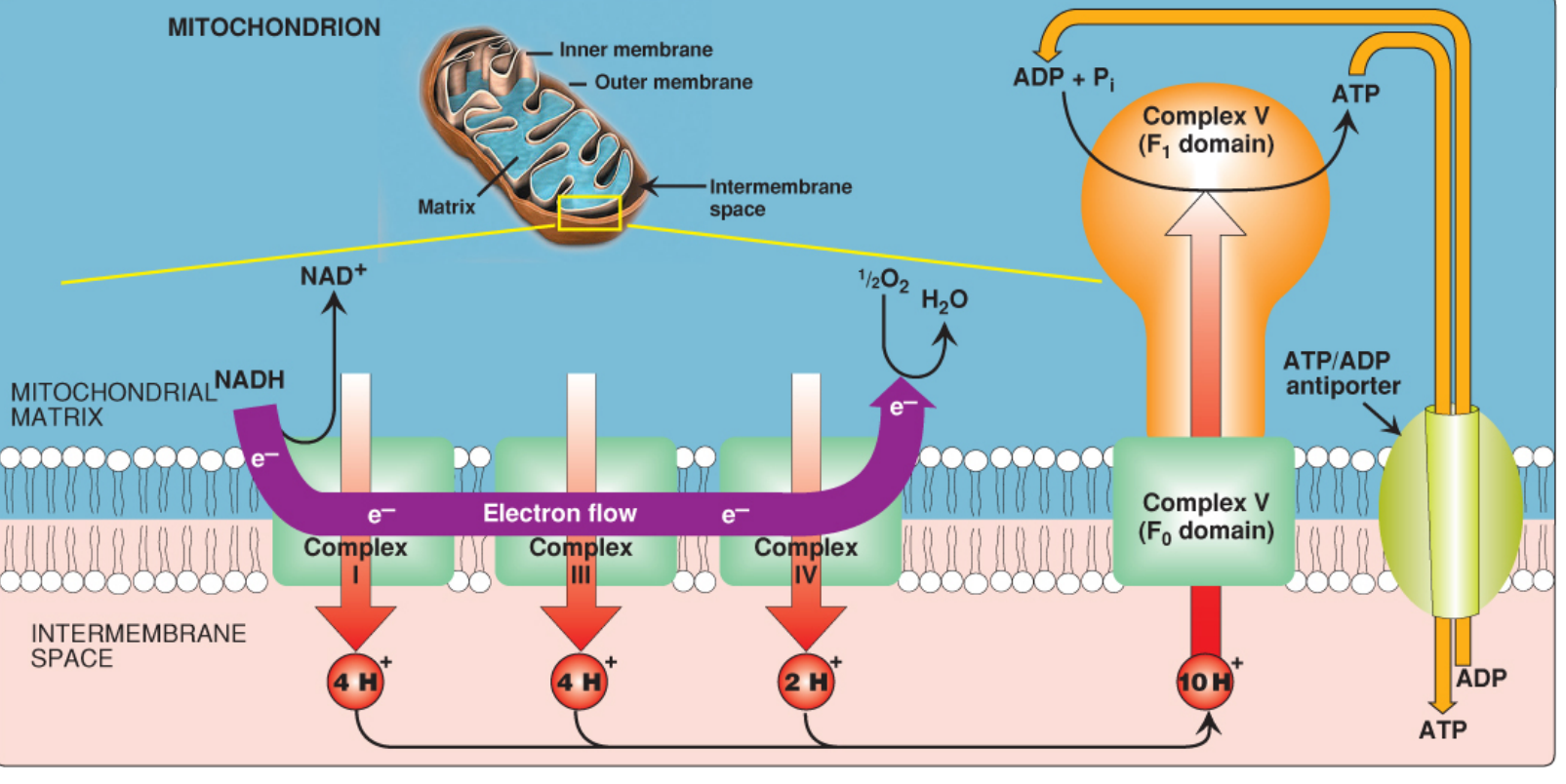
ATP Formation — Chemiosmotic Hypothesis (Mitchell Hypothesis)
Mechanism
-
Proton pump: As electrons pass through Complexes I, III, and IV, energy released drives pumping of H⁺ from the matrix to the intermembrane space (IMS).
- 10 H⁺ pumped per NADH oxidized (4+4+2)
- 6 H⁺ pumped per FADH₂ oxidized (0+4+2, entering via CoQ)
-
Electrochemical gradient (Proton-Motive Force): This creates:
- An electrical gradient (positive charge accumulates in IMS)
- A pH gradient (IMS is more acidic than matrix)
- Together = proton-motive force (~220 mV)
-
ATP synthase (Complex V): H⁺ re-enter the matrix through the F₀ channel, driving rotation of the c-ring. This rotational energy is transmitted to the F₁ domain, where conformational changes in the three β subunits cyclically:
- Bind ADP + Pᵢ
- Catalyze phosphorylation (ATP synthesis)
- Release ATP
- One full rotation = 3 ATP; ~3–4 H⁺ needed per ATP
Binding Change Mechanism (Boyer)
The three β subunits exist in three conformational states (open "O", loose "L", tight "T") that rotate with each H⁺ flux. ADP + Pᵢ bind at the open site; ATP is spontaneously synthesized at the tight site; and pre-formed ATP is released as the site becomes open again.
ATP Synthesis Sites
| Site | ATP Yield per Mole |
|---|---|
| NADH → Complex V (via I, III, IV) | ~2.5 ATP |
| FADH₂ → Complex V (via II, III, IV) | ~1.5 ATP |
| Substrate-level phosphorylation (TCA — succinyl-CoA synthetase) | 1 GTP/ATP |
| Substrate-level phosphorylation (Glycolysis — PGK, PK) | 2 ATP net |
Total ATP from complete glucose oxidation:
- Glycolysis: ~2 ATP (net) + 2 NADH (cytosolic)
- Pyruvate decarboxylation: 2 NADH
- TCA cycle: 6 NADH + 2 FADH₂ + 2 GTP
- Grand total: ~30–32 ATP per glucose
Diagram 3 — Full ETC with ROS Generation
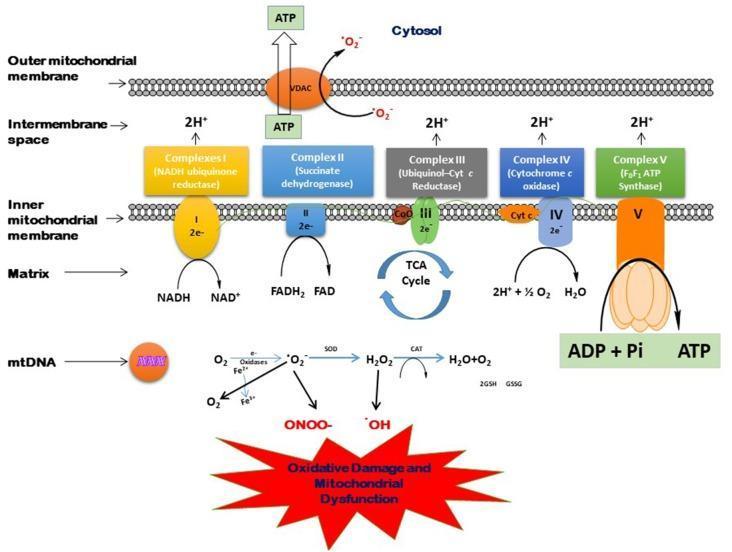
Inhibitors of the ETC
A. ETC Electron Transport Inhibitors (Block specific complexes)
| Inhibitor | Site of Action | Mechanism | Clinical/Toxicological Significance |
|---|---|---|---|
| Rotenone | Complex I | Blocks NADH → CoQ electron transfer | Pesticide/insecticide; used in Parkinson's research |
| Amobarbital (Amytal) | Complex I | Blocks FMN/Fe-S region | Barbiturate toxicity |
| TTFA (Thenoyltrifluoroacetone) | Complex II | Blocks Fe-S centers | Experimental inhibitor |
| Antimycin A | Complex III | Blocks Cyt b → Cyt c₁ (inhibits Q cycle) | Fungicide; fish poison |
| BAL (Dimercaprol) | Complex III | Chelates Fe-S centers | Also used as chelation therapy |
| CO (Carbon monoxide) | Complex IV | Binds Cyt a₃, blocks O₂ binding | Lethal poisoning; binds with high affinity |
| Cyanide (CN⁻) | Complex IV | Binds Fe³⁺ of Cyt a₃ | Used in chemical warfare; treated with hydroxocobalamin/thiosulfate |
| Hydrogen sulfide (H₂S) | Complex IV | Similar to CN⁻ | Sewer gas poisoning |
| Azide (N₃⁻) | Complex IV | Binds Cyt a₃ | Laboratory ETC inhibitor |
Effect of ETC inhibitors: Electron flow stops → H⁺ pumping stops → proton-motive force collapses → ATP synthesis stops → NADH and FADH₂ accumulate → O₂ consumption decreases.
B. ATP Synthase (Phosphorylation) Inhibitors
| Inhibitor | Mechanism |
|---|---|
| Oligomycin | Binds F₀ domain; blocks H⁺ channel → stops proton re-entry → electron transport also stops (respiratory control) |
| Aurovertin | Binds F₁ domain; inhibits ATP synthesis |
| Dicyclohexylcarbodiimide (DCCD) | Reacts covalently with F₀; blocks proton channel |
C. Uncouplers of Oxidative Phosphorylation
Uncouplers dissipate the proton gradient without going through ATP synthase. Electron transport continues (or accelerates), but energy is released as heat instead of ATP.
| Uncoupler | Type | Mechanism |
|---|---|---|
| 2,4-Dinitrophenol (DNP) | Synthetic (lipophilic ionophore) | Shuttles H⁺ across inner membrane, bypassing F₀ |
| CCCP (Carbonyl cyanide m-chlorophenylhydrazone) | Synthetic | Protonophore; causes uncoupling |
| Thermogenin / UCP1 | Endogenous (brown fat) | H⁺ channel in inner mitochondrial membrane; non-shivering thermogenesis in neonates; activated by cold via catecholamines |
| UCP2, UCP3 | Endogenous (other tissues) | Role in heat production, regulation of ROS |
| Aspirin (salicylates) — high dose | Drug | Uncouples OXPHOS; mechanism of fever in overdose |
| Long-chain fatty acids | Endogenous | Act as protonophores; can be returned to normal by albumin |
Key feature of uncouplers: O₂ consumption increases (electron transport uninhibited) but ATP synthesis decreases. Body temperature rises (energy released as heat). Weight loss occurs as more fuel is burned without ATP production — the historical basis for DNP's brief use as a diet drug (abandoned due to fatal hyperthermia).
Respiratory Control
In tightly coupled mitochondria, electron transport rate is controlled by ADP availability:
- High ADP → faster electron transport, more O₂ consumption, more ATP synthesis
- Low ADP → electron transport slows (rate-limiting step is H⁺ re-entry through ATP synthase)
- Oligomycin locks mitochondria in a state of low electron transport despite adequate substrate
Summary: Sites of Action
NADH → [Complex I] → CoQ ← [Complex II] ← FADH₂
↓
[Complex III] → Cyt c
↓
[Complex IV] → H₂O
↓
H⁺ gradient (proton-motive force)
↓
[Complex V / ATP Synthase]
↓
ATP
Inhibitors: Uncouplers:
I — Rotenone DNP, Thermogenin (UCP1)
III — Antimycin A Salicylates (high dose)
IV — CN⁻, CO, N₃⁻
V — Oligomycin
Sources:
- Lippincott Illustrated Reviews: Biochemistry, 8th ed. pp. 224–243
- Basic Medical Biochemistry: A Clinical Approach, 6e. pp. 860–878
This is a shared conversation. Sign in to Orris to start your own chat.