What is maternal cell free DNA and it's uses in pregnancy
Reading File
Finding Sources
Finding Sources
Searching PubMed
"cell-free DNA" AND "prenatal screening"
Reading File
Reading File
Reading File
Reading File
Reading File
Loading Image
I now have comprehensive information from multiple authoritative textbooks. Let me compile a thorough answer.
Maternal Cell-Free DNA (cfDNA) in Pregnancy
What Is It?
Cell-free DNA (cfDNA) refers to small fragments of DNA that circulate freely in the bloodstream, outside of any cell. All individuals have cfDNA in their plasma, but during pregnancy, maternal blood also contains cell-free fetal DNA (cffDNA) — fragments derived from apoptosis of trophoblast cells of the placenta, which carry the same genome as the fetus.
Key physical characteristics:
- Fragments are short (typically <450 base pairs, many <100 bp)
- Fetal fragments are generally shorter than maternal fragments — this size difference can be exploited analytically
- Undergoes rapid turnover: mean half-life of ~16 minutes at term
- Detectable in maternal plasma from as early as 18 days after embryo transfer
- Becomes undetectable ~2 hours after delivery
— Creasy & Resnik's Maternal-Fetal Medicine, p. 657
The Fetal Fraction
The proportion of cfDNA in maternal blood that is fetal in origin is called the fetal fraction. After 10 weeks' gestation:
- Fetal fraction is approximately 5%–20% of total circulating cfDNA
- Levels increase until ~10 weeks, plateau between 10–21 weeks, then rise again in the third trimester
- A minimum fetal fraction (typically ≥4%) is required for reliable testing; below this threshold, a "no-result" may occur
— Thompson & Thompson Genetics and Genomics in Medicine, 9th ed., p. 407
How It Is Analyzed: Sequencing Methods
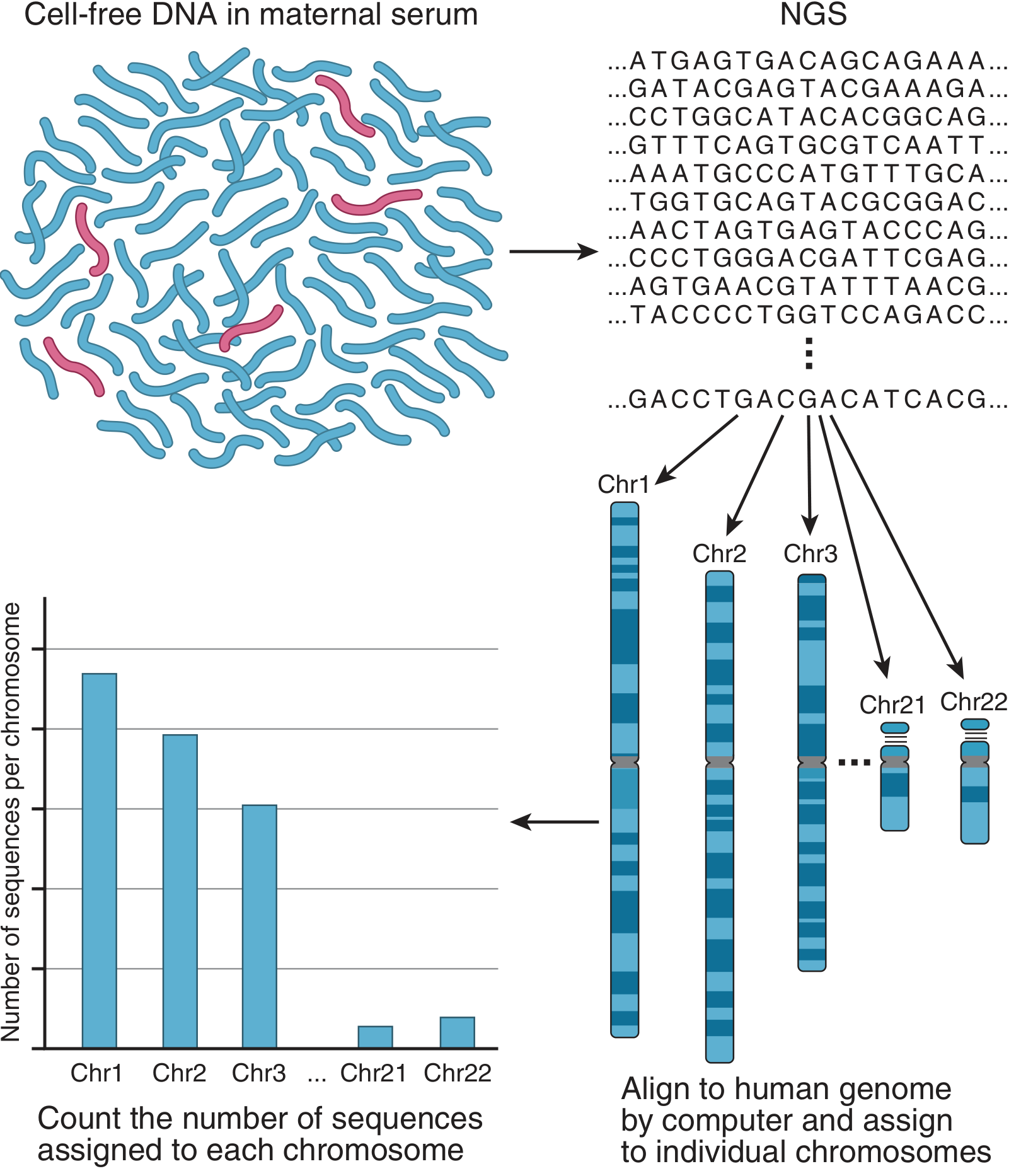
Figure: Noninvasive prenatal screening (NIPS) by cfDNA analysis. Fetal DNA fragments (red) are a minority within maternal DNA (blue). Millions of fragments are sequenced, mapped to their chromosome of origin, and counted. An excess from chromosome 21 indicates trisomy 21.
Three main approaches are used:
| Method | Principle |
|---|---|
| Massively Parallel Shotgun Sequencing (MPSS) | All circulating DNA sequenced; fragments aligned to genome; relative chromosome representation counted |
| Targeted Massively Parallel Sequencing | Selects specific chromosomal loci (e.g., chromosomes 21, 18, 13) for focused analysis |
| SNP-based analysis | Uses single-nucleotide polymorphisms to distinguish maternal vs. fetal DNA and detect copy number changes |
In a trisomy 21 pregnancy where 10% of cfDNA is fetal, only ~1.05× more chromosome 21 fragments than expected appear — the system must sequence millions of fragments to detect this small but significant difference.
— Creasy & Resnik's, p. 658
Clinical Uses in Pregnancy
1. Screening for Common Fetal Aneuploidies (Primary Use — NIPT/NIPS)
Also called Non-Invasive Prenatal Testing (NIPT) or Non-Invasive Prenatal Screening (NIPS). Available from 9–10 weeks' gestation.
| Condition | Detection Rate | Notes |
|---|---|---|
| Trisomy 21 (Down syndrome) | ~99% sensitivity, ~99% specificity | Highest performance |
| Trisomy 18 (Edwards syndrome) | ~96–98% | High performance |
| Trisomy 13 (Patau syndrome) | ~92–98% | Good performance |
| Sex chromosome aneuploidies (45,X; 47,XXY; 47,XXX; 47,XYY) | Variable | Can be added to panel |
Key caveat: cfDNA is a screening test, not a diagnostic test. A positive result must be confirmed by CVS or amniocentesis before clinical action is taken. Negative predictive value (NPV) exceeds 99% for all common aneuploidies.
— Thompson & Thompson, p. 407–408; Creasy & Resnik's, p. 657
2. Screening for Microdeletion Syndromes
Some commercial labs extend cfDNA panels to include:
- 22q11.2 deletion (DiGeorge syndrome) — most common microdeletion
- 1p36 deletion syndrome
- Prader-Willi / Angelman syndromes (15q deletion)
- Wolf-Hirschhorn syndrome (4p deletion)
- Cri du chat syndrome (5p deletion)
The positive predictive value for microdeletions is lower than for common trisomies due to their rarity, so current US guidelines do not routinely recommend cfDNA for microdeletion screening outside of research contexts.
— Creasy & Resnik's, p. 660
3. Fetal Sex Determination
Detection of Y-chromosome sequences (SRY gene) in maternal plasma reliably identifies a male fetus. This is useful in:
- Pregnancies at risk for X-linked disorders (e.g., Duchenne muscular dystrophy, hemophilia)
- Congenital adrenal hyperplasia (to guide dexamethasone treatment decisions in female fetuses only)
4. Fetal Rhesus (RhD) Blood Group Genotyping
In RhD-negative mothers, cfDNA can determine whether the fetus is RhD-positive or -negative. This allows targeted anti-D (Rh immunoglobulin) prophylaxis — avoiding unnecessary treatment in mothers carrying RhD-negative fetuses. Widely used in several European countries as a public health intervention.
5. Single-Gene Disorder Testing
cfDNA assays are clinically available (in high-risk pregnancies) for a growing number of conditions:
| Clinically Available |
|---|
| Achondroplasia |
| Apert syndrome |
| Congenital adrenal hyperplasia |
| Cystic fibrosis |
| Duchenne/Becker muscular dystrophy |
| Spinal muscular atrophy |
| Blood group genotyping (RHD/RHCE; Kell) |
| Torsion dystonia |
| Selected familial known mutations |
Research is ongoing for hemoglobinopathies (sickle cell, thalassemias), Fraser syndrome, and whole-genome fetal sequencing.
— Thompson & Thompson, Table 18.5, p. 408
6. Whole-Genome Copy Number Analysis
Some providers offer genome-wide analysis of copy number variants (CNVs) — gains and losses across all chromosomes — though this remains less validated than targeted testing.
Limitations and Counseling Points
Per Box 30.3 of Creasy & Resnik's, key pre-test counseling points include:
- cfDNA has the highest detection for trisomy 21, but false-positive and false-negative results do occur
- A negative result does not rule out all chromosomal abnormalities or birth defects
- cfDNA does not screen for open neural tube defects — maternal serum AFP (MSAFP) or detailed ultrasound is still needed
- cfDNA detects only ~50% of cytogenetic abnormalities that would be found by karyotype on CVS/amniocentesis
- In rare cases, cfDNA can reveal an underlying chromosomal abnormality or malignancy in the mother (incidental finding)
- If a structural anomaly is seen on ultrasound, chromosomal microarray via CVS/amniocentesis is recommended over cfDNA
Recent Evidence (2024–2025)
Two recent systematic reviews are relevant:
-
Low fetal fraction and adverse pregnancy outcomes — a 2024 meta-analysis (PMID: 39008086) found that low fetal fraction on cfDNA testing is associated with adverse pregnancy outcomes beyond failed test results, suggesting it may carry prognostic value.
-
Screening for fetal de novo / paternally inherited single nucleotide variants — a 2025 systematic review (PMID: 40411478) assessed feasibility of cfDNA-based SNV detection, pointing toward expanded future applications of NIPT beyond chromosomal aneuploidies.
In summary, maternal cfDNA has transformed prenatal screening by enabling accurate, non-invasive detection of fetal chromosomal conditions from as early as 9 weeks. Its primary, validated use is aneuploidy screening (NIPT/NIPS), with expanding applications in single-gene disorders, RhD genotyping, and potentially whole-genome fetal assessment. It remains a screening — not diagnostic — tool, and positive results always require invasive confirmation.
Type of twins and the embryological leading to those types
Finding Sources
Finding Sources
Finding Sources
Finding Sources
Reading File
Reading File
Reading File
Reading File
Reading File
Loading Image
Reading File
I now have comprehensive information from multiple authoritative embryology and obstetrics textbooks. Let me now compile a thorough, well-organized answer.
Types of Twins and Their Embryological Basis
Overview
Twins arise by two fundamentally different mechanisms:
| Dizygotic (DZ) | Monozygotic (MZ) | |
|---|---|---|
| Common name | Fraternal / non-identical | Identical |
| Origin | Two separate zygotes | One zygote, split after fertilization |
| Genetics | ~50% shared (like siblings) | Genetically identical |
| Sex concordance | May differ | Always same sex (usually) |
| Frequency | ~90% of all twins | ~10% of all twins; rate 3–4/1,000 |
| Chorionicity | Always dichorionic | Depends on timing of split |
1. Dizygotic (Fraternal) Twins
Embryological Mechanism
DZ twins result from simultaneous ovulation and fertilization of two separate oocytes by two different spermatozoa. Each zygote implants separately in the uterus and develops its own placenta, amnion, and chorionic sac — making them always dichorionic–diamniotic (DCDA).
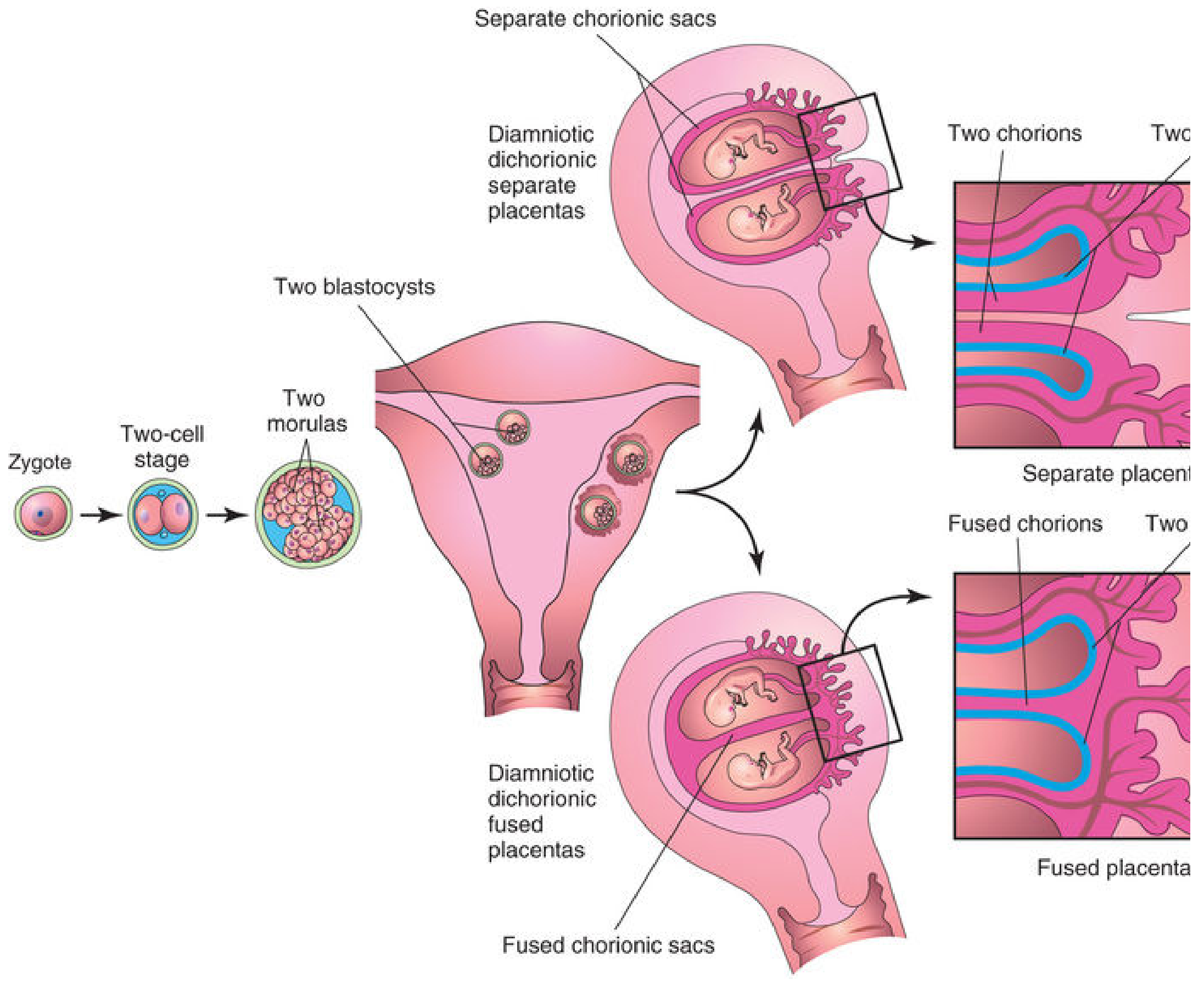
Dizygotic twinning: two zygotes → two morulas → two blastocysts → separate implantations. Placentas may be completely separate or fused if implantation sites are close together.
Key facts:
- When the two implantation sites are adjacent, the placentas and chorionic walls may fuse — but they remain two distinct placentas histologically
- In rare cases of fused placentas, red blood cells can be exchanged → erythrocyte mosaicism
- In a male–female DZ pair, unlike cattle (freemartin effect), masculinization of the female does NOT occur in humans
- Incidence is influenced by maternal age (peaks ~35 years), ART/fertility drugs, and ethnicity:
- ~1 in 500 in Asians
- ~1 in 125 in White Europeans
- As high as ~1 in 20 in some sub-Saharan African populations
- Shows hereditary tendency: recurrence ~3× general population; the maternal genotype affects DZ twinning rate; paternal genotype does not
— The Developing Human, p. 399; Langman's Medical Embryology, p. 8.18
2. Monozygotic (Identical) Twins
MZ twins originate from a single fertilized ovum that splits. The timing of splitting is the critical embryological variable — it determines the chorionicity and amnionicity of the twins, and therefore their clinical risk profile.
The Four Subtypes Based on Timing of Splitting
Type 1: Dichorionic–Diamniotic (DCDA) MZ Twins
Splitting at 2-cell to morula stage (~days 1–3 post-fertilization; before inner cell mass forms)
- Two separate blastocysts form and implant independently
- Each embryo develops its own amnion, chorion, and placenta
- Placentas may be separate or fused (secondary fusion)
- Accounts for ~35% of MZ twins
- Clinically indistinguishable from DZ twins by membrane analysis alone — only genetic testing confirms MZ identity
- Lowest risk subtype
Type 2: Monochorionic–Diamniotic (MCDA) MZ Twins ← Most Common MZ Type
Splitting at early blastocyst stage (~days 4–8; inner cell mass splits within the same blastocyst cavity)
- Two embryonic primordia form within one chorion but develop separate amniotic sacs
- Twins share a single placenta and chorion but have two amnions
- Accounts for ~65% of MZ twins (the most common arrangement)
- Higher risk: shared placenta predisposes to Twin-to-Twin Transfusion Syndrome (TTTS), affecting 10–15% of MCDA pairs
Type 3: Monochorionic–Monoamniotic (MCMA) MZ Twins
Splitting at bilaminar germ disc stage (~days 8–13; just before primitive streak appears)
- Division is very late — both embryos share one chorion, one amnion, and one placenta
- Occurs in ~1% of MZ gestations
- Highest spontaneous risk: umbilical cord entanglement is almost universal; historically carried >50% perinatal mortality; modern intensive surveillance has reduced this significantly
- Diagnosed sonographically by absence of inter-twin membrane; entangled cords confirmed with colour Doppler
- Management: intensive fetal surveillance from viability; elective delivery typically at 32–34 weeks
Type 4: Conjoined Twins
Incomplete splitting after day 13–15 (embryonic disc stage), or fusion of adjacent embryonic discs
- Splitting fails to complete → physical union at a shared anatomic site
- Estimated frequency: ~1.5 in 100,000 births (some sources cite 1 in 50,000–100,000)
- More common in females than males
- Always monochorionic–monoamniotic
Classification by site of fusion:
| Type | Fusion Site | Frequency | Prognosis |
|---|---|---|---|
| Thoracopagus | Anterior thorax; face each other | ~75% | Often poor (75% share hearts) |
| Omphalopagus (xiphopagus) | Anterior abdominal wall, liver | Rare subtype of thoracopagus | Better (hearts rarely joined) |
| Pygopagus | Sacrum; face away | ~20% | Usually good after separation |
| Ischiopagus | Bony pelvis | ~5% | Good prognosis; lower spine often abnormal |
| Craniopagus | Skull, meninges, vasculature | ~1% | Depends on vascular fusion |
A rare variant: heteropagus (parasitic) twins — one severely defective twin is wholly dependent on the cardiovascular system of the intact twin (incidence ~1 in 1,000,000).
— Creasy & Resnik's Maternal-Fetal Medicine, p. 858; The Developing Human, p. 404
Summary: MZ Twin Types by Timing of Splitting
| Day of Splitting | Stage | Chorionicity | Amnionicity | Frequency (of MZ) |
|---|---|---|---|---|
| Days 1–3 | 2-cell → morula | Dichorionic | Diamniotic | ~35% |
| Days 4–8 | Early blastocyst (ICM splits) | Monochorionic | Diamniotic | ~65% |
| Days 8–13 | Bilaminar disc (late blastocyst) | Monochorionic | Monoamniotic | ~1% |
| Day 13–15+ | Embryonic disc (incomplete split) | Monochorionic | Monoamniotic | Rare (conjoined) |
The later the split, the more structures are shared — and the higher the clinical risk.
Special Concepts
Twin Transfusion Syndrome (TTTS)
Occurs in 10–15% of MCDA twins. Unidirectional arteriovenous placental anastomoses shunt blood from the donor twin (pale, anemic, small, oligohydramnios) to the recipient twin (plethoric, polycythemia, large, polyhydramnios). Severe cases can cause death from anemia in the donor and congestive heart failure in the recipient. Treatment: fetoscopic laser coagulation of the anastomoses.
— The Developing Human, p. 402
Discordance in MZ Twins
MZ twins may be discordant for birth defects or genetic disorders despite identical genomes, due to:
- Vascular abnormalities during development (e.g., TTTS)
- Differences in X-chromosome inactivation
- Post-zygotic somatic mutations
- Epigenetic differences
Superfecundation
Fertilization of two oocytes at different times (or by different fathers — confirmed in humans by genetic markers). A rare mechanism contributing to DZ twinning.
Zygosity Determination
Clinical examination of placental membranes can be misleading — 35% of MZ twins are DCDA and may be misclassified as DZ. Definitive determination requires DNA marker analysis (molecular diagnosis).
— The Developing Human, p. 401; Langman's, p. 8.19
This is a shared conversation. Sign in to Orris to start your own chat.