hemoglobin electrophoresis
Reading File
Finding Sources
Finding Sources
Searching Images
hemoglobin electrophoresis pattern sickle cell
Reading File
Reading File
Loading Image
Loading Image
Reading File
I now have comprehensive information. Let me compile a thorough answer.
Hemoglobin Electrophoresis
Principle
Hemoglobin electrophoresis separates hemoglobin variants based on molecular charge and size. Different globin chain mutations alter the surface charge of the tetramer, causing variants to migrate at different rates through the gel toward the anode (+) or cathode (−).
Methods
1. Alkaline Electrophoresis (pH 8.6) — Cellulose Acetate
The standard first-line method. Migration order from cathode → anode:
Origin → Carbonic anhydrase → C / A2 / E / O → S / D / G / Lepore → F → A / M → Barts / H (fastest)
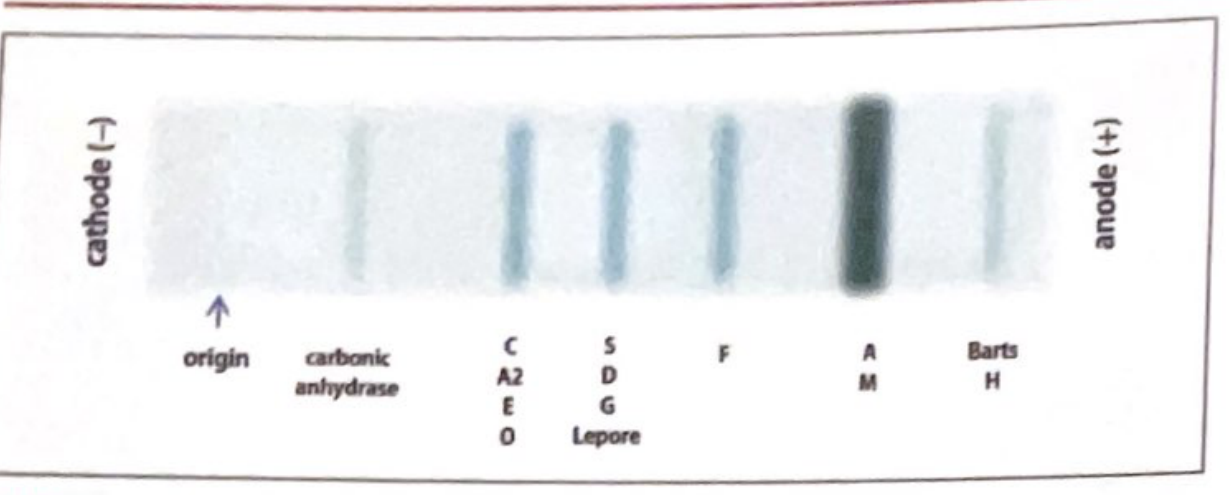
Normal adult: >97% HbA + <3% HbA2 (in the C band) + nothing else.
Key limitation: Cannot separate HbS from HbD, HbG, or Hb-Lepore. A band in the S region must be confirmed with a sickle screen (metabisulfite or solubility test). Fast hemoglobins (HbH, Hb Barts) can be mimicked by hyperbilirubinemia.
2. Acid Electrophoresis (pH 6.2) — Citrate Agar
Used as a confirmatory/complementary test when alkaline results are ambiguous.
Migration order from cathode → anode:
F → A / A2 / D / G / E → Origin → S / C-Harlem → C
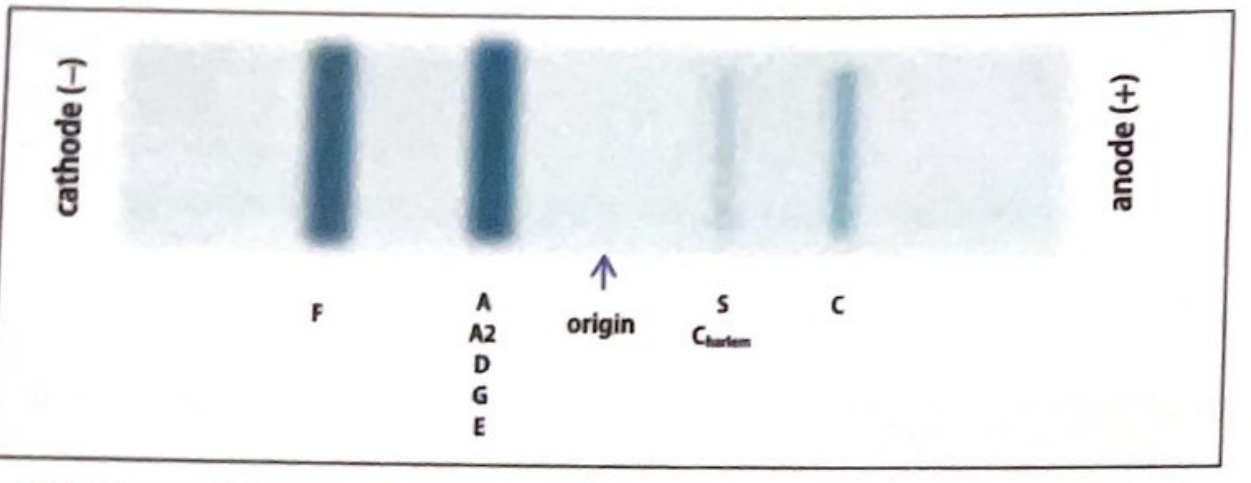
Key point: At acid pH, HbS and HbC now run separately (unlike some co-migrations at alkaline pH), helping confirm HbS. However, it still cannot separate HbD from HbG/Hb-Lepore, or HbE from HbO-Arab.
Important: Electrophoresis does not permit accurate quantitation of HbA2 or HbF — HPLC is needed for that.
3. HPLC (High-Performance Liquid Chromatography)
The most accurate quantitative method. Molecules elute at different characteristic retention times, allowing both identification and precise quantification of HbA2 and HbF.
Limitations:
- HbE and HbA2 have similar retention times (not easily separated)
- HbC and HbO-Arab have similar retention times
- Bilirubin co-elutes with Hb Barts
4. Isoelectric Focusing (IEF) / Capillary Electrophoresis
- IEF is a component of mandatory newborn screening in many states
- Capillary electrophoresis can accurately quantify low HbA2 levels and separately quantify HbE levels (though HbC still runs with HbA2)
5. Molecular Methods (PCR/Gene Sequencing)
~1–2% of variant hemoglobins detected by HPLC or gel electrophoresis cannot be definitively identified — PCR sequencing of the globin gene characterizes the exact genotype.
Neonatal Hemoglobin Electrophoresis Patterns
Variants are reported in order of decreasing abundance (e.g., "FA" = more HbF than HbA):
| Pattern | Interpretation |
|---|---|
| FA | Normal newborn (HbF predominant, HbA present) |
| FAS | Sickle cell trait (benign) |
| FS | Sickle cell disease (SS) or Sβ⁰-thalassemia — clinically significant |
| FSC | HbSC disease — clinically significant sickle disorder |
| FSA | Sβ⁺-thalassemia — clinically significant sickling disorder |
| FC | Homozygous HbC (C/C) — mild hematologic disorder |
| FAC | HbC trait (benign) |
| F only | May be β-thalassemia major, hereditary persistence of HbF (HPFH), or delayed HbA switch — confirm with repeat |
| AF | May indicate prior blood transfusion — repeat at 4 months |
| FAV / FV | Anomalous band present — requires further workup |
All positive sickle solubility tests (Sickledex) must be confirmed with electrophoresis or IEF.
Interpreting Adult Patterns for Common Hemoglobinopathies
| Condition | Alkaline Gel Findings |
|---|---|
| Normal | HbA >97%, HbA2 <3% |
| Sickle cell trait (AS) | HbA + HbS band (HbA predominant) |
| Sickle cell disease (SS) | HbS only (no HbA), elevated HbF |
| HbSC disease | HbS + HbC (equal bands), no HbA |
| β-thalassemia minor | Thalassemic indices + elevated HbA2 >3.5% |
| α-thalassemia | Thalassemic indices + normal HbA2 |
| HbC trait | HbA + HbC band |
| HbE trait | HbA + HbE (in C/A2 band on alkaline) |
Clinical Indications
- Newborn screening (mandatory in most U.S. states)
- Diagnosis of sickle cell disease/trait
- Evaluation of thalassemia
- Workup of unexplained hemolytic anemia, microcytosis, or elevated HbF
- Thrombophilia evaluation (sickle hemoglobins cause vascular occlusion)
- Prenatal carrier screening (combined with MCV)
- Confirmation of positive sickle solubility tests
Sources: Quick Compendium of Clinical Pathology, 5th ed.; The Harriet Lane Handbook, 23rd ed.
neoplasia
Asking for Preferences
Finding Sources
Finding Sources
Reading File
Finding Sources
Finding Sources
Finding Sources
Reading File
Finding Sources
Reading File
Searching Images
benign vs malignant tumor characteristics differentiation anaplasia histology
Reading File
Reading File
I now have comprehensive material from Robbins. Let me compile the full answer.
Neoplasia — General Concepts, Definitions & Classification
Definition
Neoplasia ("new growth") refers to an abnormal mass of tissue in which growth exceeds and is uncoordinated with that of normal tissues, and which persists after cessation of the stimuli that initiated it. Neoplastic cells are said to be transformed — they replicate incessantly due to resistance to normal regulatory controls, enjoying a degree of autonomy while still depending on the host for nutrition and blood supply.
In common usage, a neoplasm is called a tumor, and the study of tumors is oncology (from oncos, tumor).
Two Basic Components of All Tumors
Every tumor — benign or malignant — has two components:
- Parenchyma — the transformed/neoplastic cells (determines biologic behavior, gives the tumor its name)
- Stroma — host-derived supporting tissue: connective tissue, inflammatory cells, blood vessels (essential for growth and nutrition; carries on a two-way conversation with tumor cells)
Nomenclature
Benign Tumors
Named by attaching the suffix -oma to the cell type of origin:
| Special terms | Meaning |
|---|---|
| Adenoma | Benign epithelial neoplasm producing gland-like structures (or from glandular epithelium) |
| Papilloma | Benign epithelial neoplasm forming finger-like fronds |
| Polyp | Mass projecting above a mucosal surface (macroscopically visible; can be benign or rarely malignant) |
| Cystadenoma | Hollow cystic benign mass (common in ovary) |
Malignant Tumors
| Origin | Malignant term |
|---|---|
| Epithelium (any germ layer) | Carcinoma |
| — glandular pattern | Adenocarcinoma |
| — squamous pattern | Squamous cell carcinoma |
| Mesenchymal/solid tissue | Sarcoma (e.g., liposarcoma, chondrosarcoma) |
| Blood cells | Leukemia |
| Lymphoid tissue | Lymphoma |
Nomenclature Table (Selected Examples)
| Tissue | Benign | Malignant |
|---|---|---|
| Fibrous tissue | Fibroma | Fibrosarcoma |
| Fat | Lipoma | Liposarcoma |
| Cartilage | Chondroma | Chondrosarcoma |
| Bone | Osteoma | Osteosarcoma |
| Blood vessels | Hemangioma | Angiosarcoma |
| Smooth muscle | Leiomyoma | Leiomyosarcoma |
| Striated muscle | Rhabdomyoma | Rhabdomyosarcoma |
| Squamous epithelium | Squamous papilloma | Squamous cell carcinoma |
| Glandular epithelium | Adenoma | Adenocarcinoma |
| Melanocytes | Nevus | Melanoma |
Important exceptions: Some malignant tumors have confusing "-oma" names — lymphoma, melanoma, mesothelioma, hepatoma (hepatocellular carcinoma) are all malignant despite the suffix.
Mixed Tumors
When the progenitor cell can differentiate along more than one lineage:
- Pleomorphic adenoma (salivary gland) — epithelial + fibromyxoid stroma ± cartilage/bone
- Teratoma — contains tissues from >1 (sometimes all 3) germ cell layers; originates from totipotent germ cells (ovary, testis, midline rests)
Characteristics: Benign vs. Malignant
| Feature | Benign | Malignant |
|---|---|---|
| Differentiation | Well-differentiated; closely resembles tissue of origin | Poorly to undifferentiated (anaplastic); may bear little resemblance to origin |
| Growth rate | Usually slow | Often rapid (variable) |
| Mitotic figures | Rare; normal morphology | Frequent; may be atypical/abnormal |
| Nuclear morphology | Normal N:C ratio, fine chromatin | High N:C ratio, hyperchromatic nuclei, prominent nucleoli, pleomorphism |
| Borders | Well-circumscribed, often encapsulated | Poorly circumscribed, no capsule, irregular borders |
| Local invasion | Non-invasive; pushes rather than infiltrates | Invasive — infiltrates surrounding normal tissue; destroys basement membrane |
| Metastasis | None | Present (the defining feature of malignancy) |
| Recurrence | Rare after excision | Common |
| Effect on host | Usually local/mechanical | Systemic (cachexia, paraneoplastic syndromes), often fatal if untreated |
Key Concepts
Differentiation and Anaplasia
- Differentiation = degree to which neoplastic cells resemble their normal counterparts morphologically and functionally
- Anaplasia = loss of differentiation; hallmark of malignancy
- Features: pleomorphism (variation in cell/nuclear size and shape), abnormal nuclear morphology (hyperchromasia, coarse chromatin, large nucleoli), high mitotic index, atypical mitoses, tumor giant cells, loss of polarity
Local Invasion
- Benign tumors remain localized within a fibrous capsule
- Malignant tumors lack a true capsule; they infiltrate surrounding tissue by secreting proteases (MMPs) that degrade basement membrane and extracellular matrix
Metastasis
- The single most reliable criterion distinguishing malignant from benign
- Routes: lymphatic (carcinomas favor), hematogenous (sarcomas favor), seeding of body cavities
- Not all cancers metastasize equally (e.g., basal cell carcinoma rarely metastasizes despite being malignant)
Dysplasia
- A pre-neoplastic state: disordered cellular growth with loss of uniformity and architectural orientation, but without invasion
- When severe and involving the full thickness of an epithelium = carcinoma in situ (CIS)
- Dysplasia does not necessarily progress to cancer, but represents increased risk
Tumor vs. Host Relationship
| Feature | Detail |
|---|---|
| Dependence on host | All tumors need host vasculature (angiogenesis) and nutrition |
| Hormone-dependent tumors | Breast, prostate, endometrium — require hormonal support (exploitable therapeutically) |
| Stroma importance | Stroma provides blood supply; tumor–stromal crosstalk shapes growth and invasion |
Source: Robbins & Kumar Basic Pathology — Chapter 6: Neoplasia
Describe the pathogenesis of neoplasia
Finding Sources
Finding Sources
Finding Sources
Finding Sources
Reading File
Reading File
Reading File
Reading File
You've run out of free credits. Sign up for a free account to continue using Orris.
This is a shared conversation. Sign in to Orris to start your own chat.