Explain me structure and functions of proteins for a 1st year mbbs student
Reading File
Finding Sources
Finding Sources
Finding Sources
Reading File
Reading File
Finding Sources
Finding Sources
Finding Sources
Finding Sources
Reading File
Reading File
Reading File
Loading Image
I have all the material I need. Now let me compile a comprehensive, student-friendly explanation.
Structure and Functions of Proteins
For 1st Year MBBS — Biochemistry
What Are Proteins?
Proteins are the most functionally diverse biomolecules in the body. They are polymers of amino acids joined by peptide bonds, and every protein's unique characteristics are determined by its linear sequence of amino acids (its primary structure). From that sequence flows everything — how it folds, how it interacts with other molecules, and what function it performs.
1. Amino Acids — The Building Blocks
All 20 amino acids used for protein synthesis share the same general structure:
- A central α-carbon
- An amino group (–NH₂)
- A carboxyl group (–COOH)
- A hydrogen atom
- A unique side chain (R group) that gives each amino acid its identity
At physiologic pH (~7.4), the amino group is positively charged and the carboxylate is negatively charged — this double-charged form is called a zwitterion. All amino acids in mammalian proteins are in the L-configuration.
All amino acids in human proteins are L-α-amino acids (except glycine, which is neither D nor L because its α-carbon has two identical hydrogen substituents).
Amino acids are linked by peptide bonds — a covalent bond between the carboxyl group of one amino acid and the amino group of the next, with loss of water (condensation reaction).
2. The Four Levels of Protein Structure
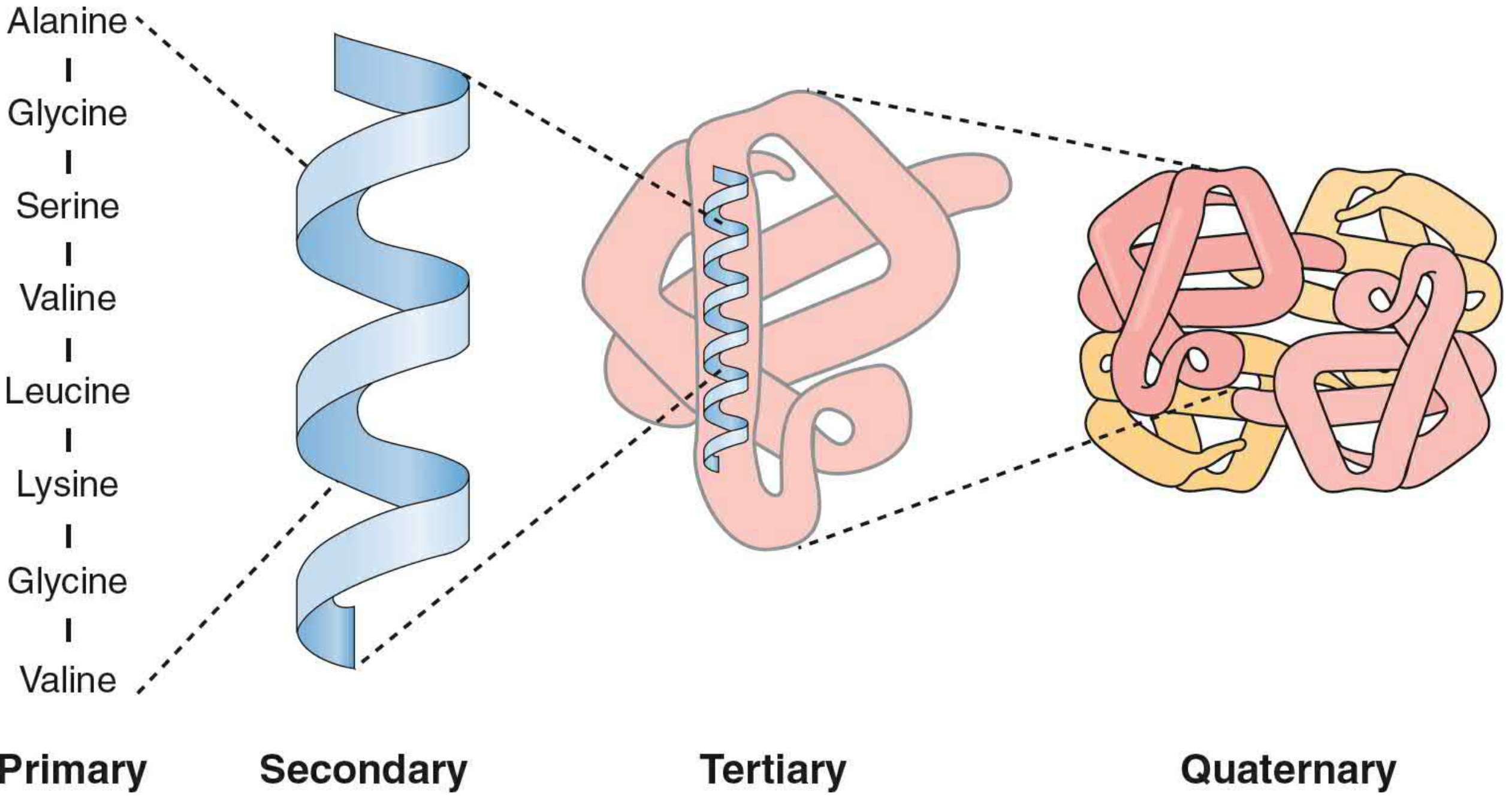
🔵 Primary Structure
The linear sequence of amino acids in the polypeptide chain. This is dictated by the genetic code (DNA → mRNA → protein). Even a single amino acid change can have profound consequences — for example, in sickle cell anemia, a single mutation changes glutamic acid → valine in the β-chain of hemoglobin, causing the disease.
Primary structure determines all higher levels of structure.
🔵 Secondary Structure
Local, regularly repeating folding patterns in segments of the polypeptide chain, stabilized by hydrogen bonds between backbone atoms (not side chains).
Two major types:
| Structure | Description | Stabilized by |
|---|---|---|
| α-Helix | Right-handed coil; 3.6 amino acids per turn; peptide backbone forms the core, R-groups point outward | H-bonds between C=O of residue n and N–H of residue n+4 |
| β-Sheet | Extended strands aligned side-by-side; can be parallel or antiparallel | H-bonds between strands |
Other secondary structures include β-turns (reverse turns) and loops/coils (irregular regions).
Key concept: The peptide bond has partial double-bond character — it is rigid and planar. Only the φ (phi) and ψ (psi) angles around the α-carbon can rotate. Allowed combinations of φ and ψ are plotted on a Ramachandran plot, which predicts permissible secondary structures.
🔵 Tertiary Structure
The overall 3D conformation of the entire polypeptide chain — how secondary structure elements fold relative to each other. This is the functional shape of the protein.
Forces stabilizing tertiary structure:
| Force | Description |
|---|---|
| Hydrophobic interactions | Nonpolar R-groups cluster in the interior away from water (strongest driving force for globular proteins) |
| Hydrogen bonds | Between polar side chains or between side chains and water |
| Ionic bonds (salt bridges) | Between oppositely charged R-groups |
| Disulfide bonds (–S–S–) | Covalent bonds between two cysteine residues; strongest bonds |
| Van der Waals forces | Weak, transient, but numerous |
Globular proteins (e.g., enzymes, antibodies) have a hydrophobic core with polar residues on the outer surface — this makes them water-soluble.
Fibrous proteins (e.g., collagen, keratin) are extended and highly asymmetric, serving structural roles.
Domains: Large proteins can fold into independent structural units called domains. Each domain folds independently and may carry out a specific function.
🔵 Quaternary Structure
Applies to proteins made of two or more polypeptide subunits (protomers). Describes how these subunits associate in space.
Classic example: Hemoglobin — 2α + 2β subunits
- Quaternary assembly allows cooperative O₂ binding (the binding of one O₂ facilitates the next)
- Forms binding sites for complex molecules (e.g., antigen binding by immunoglobulins)
- Increases protein stability
Not all proteins have quaternary structure. Myoglobin (single chain) has no quaternary structure; hemoglobin does.
3. Protein Folding and Denaturation
Folding
- The primary structure dictates the final 3D conformation (the "native conformation")
- Newly synthesized polypeptides are assisted by chaperones (chaperonins) — proteins that prevent premature misfolding and guide the chain to its correct shape
- Protein disulfide isomerase helps form correct disulfide bonds
- Peptidylproline cis-trans isomerase catalyzes isomerization of proline peptide bonds
Denaturation
- Heat, strong acids/bases, urea, detergents, and organic solvents disrupt non-covalent forces
- The protein loses its native conformation and becomes non-functional
- Denaturation is often irreversible
- Example: cooking an egg white (albumin)
Misfolding and Disease (Important for Clinical Context!)
| Disease | Mechanism |
|---|---|
| Prion diseases (CJD, scrapie) | Misfolded PrP protein acts as template to refold normal PrP |
| Alzheimer's disease | Misfolded amyloid-β aggregates |
| Sickle cell anemia | Mutated HbS aggregates at low O₂ |
4. Classification of Proteins
By Shape
| Type | Shape | Axial Ratio | Example |
|---|---|---|---|
| Globular | Compact, spherical | <3 | Enzymes, antibodies, hemoglobin |
| Fibrous | Extended, rod/cable | >10 | Collagen, keratin, elastin |
By Composition
| Type | Contains | Example |
|---|---|---|
| Simple protein | Only amino acids | Albumin |
| Conjugated protein | Amino acids + non-protein prosthetic group | Hemoglobin (heme), glycoproteins (sugar), lipoproteins (lipid) |
By Function (most important for MBBS!)
| Function | Example |
|---|---|
| Structural | Collagen (bone, skin, tendons), keratin (hair, nails) |
| Enzymatic (catalytic) | Amylase, pepsin, DNA polymerase |
| Transport | Hemoglobin (O₂), albumin (fatty acids, drugs), transferrin (iron) |
| Hormonal | Insulin, glucagon, growth hormone |
| Receptor | Insulin receptor, acetylcholine receptor |
| Immunological (defense) | Immunoglobulins (antibodies), complement proteins |
| Contractile/Motor | Actin, myosin (muscle contraction) |
| Storage | Ferritin (iron), casein (milk protein) |
| Regulatory | Histones (gene regulation), transcription factors |
| Ion channel/Transport | Na⁺/K⁺-ATPase, aquaporins |
5. Ligand Binding
Proteins form specific binding sites for molecules called ligands (e.g., O₂, ATP, hormones). The affinity of a protein for its ligand is described by:
- Association constant (Kₐ) — higher Kₐ = tighter binding
- Dissociation constant (Kd) — lower Kd = tighter binding; Kₐ = 1/Kd
Quick Summary Table
| Level | What it is | Stabilizing Force | Example |
|---|---|---|---|
| Primary | AA sequence | Peptide (covalent) bonds | Sickle cell Hb (Val instead of Glu) |
| Secondary | α-helix, β-sheet | H-bonds (backbone) | α-helix in myoglobin |
| Tertiary | 3D shape of one chain | Hydrophobic, H-bonds, ionic, S–S, vdW | Globular enzymes |
| Quaternary | Multi-subunit assembly | Same as tertiary (between subunits) | Hemoglobin (2α + 2β) |
Clinical High-Yield Points for 1st Year
- Sickle cell anemia — Primary structure change (Glu→Val) → abnormal secondary/tertiary structure → HbS polymerizes → sickling
- Scurvy — Deficiency of Vitamin C impairs hydroxylation of proline in collagen → defective collagen tertiary structure → connective tissue failure
- Prion diseases — Abnormal tertiary structure propagates → neurodegenerative disease
- Denaturation of proteins — Basis of sterilization (heat kills bacteria by denaturing their proteins)
- Enzymes — Catalytic activity depends entirely on tertiary structure; denaturation = loss of enzyme activity
Sources: Basic Medical Biochemistry: A Clinical Approach (6e); Harper's Illustrated Biochemistry (32e)
Explain urea cycle with skeletal structures
Finding Sources
Searching Images
urea cycle skeletal structure chemical diagram
Reading File
Reading File
Loading Image
Reading File
Reading File
Loading Image
Reading File
Loading Image
This is a shared conversation. Sign in to Orris to start your own chat.