Line probe assay full details for md exam answer 15 marks
Reading File
Finding Sources
Finding Sources
Reading File
Reading File
Reading File
Reading File
Reading File
Reading File
Reading File
Finding Sources
Reading File
Finding Sources
Reading File
Finding Sources
Reading File
Finding Sources
Reading File
Reading File
Loading Image
I now have all the material needed. Let me compose a comprehensive 15-mark MD exam answer.
Line Probe Assay (LPA) — Comprehensive Answer
Definition
A Line Probe Assay (LPA) is a molecular diagnostic technique based on the reverse hybridization principle of solid-phase hybridization. Specific oligonucleotide probes are immobilized on a nitrocellulose membrane strip as parallel lines; biotinylated amplified target DNA (from the patient specimen) is then hybridized to these membrane-bound probes. A colorimetric reaction reveals the pattern of hybridization, allowing identification of organisms and detection of drug-resistance mutations. — Tietz Textbook of Laboratory Medicine, 7th Ed.
Principle
Reverse Hybridization (Core Concept)
Unlike conventional dot-blot hybridization (where target is fixed and probe is in solution), LPA reverses this: probes are fixed to the membrane and amplified target is in solution. This enables many probe sequences to be tested simultaneously on a single strip.
Step-by-Step Process
| Step | Detail |
|---|---|
| 1. DNA Extraction | Mycobacterial DNA is extracted from sputum (AFB smear-positive) or culture isolate. |
| 2. PCR Amplification | Target gene regions (rpoB, katG, inhA, gyrA, rrs, etc.) are amplified using biotinylated primers, labeling the amplicons. |
| 3. Denaturation | Double-stranded PCR products are denatured to single strands. |
| 4. Hybridization | Single-stranded amplicons hybridize with their complementary probes fixed on the membrane strip under stringent conditions (allows only perfect-match binding). |
| 5. Stringent Wash | Unbound or partially-bound amplicons are removed. |
| 6. Detection | Streptavidin-conjugated alkaline phosphatase binds to the biotin on the hybridized amplicon. A chromogenic substrate (NBT/BCIP) is added → purple/dark band appears at positions with hybridization. |
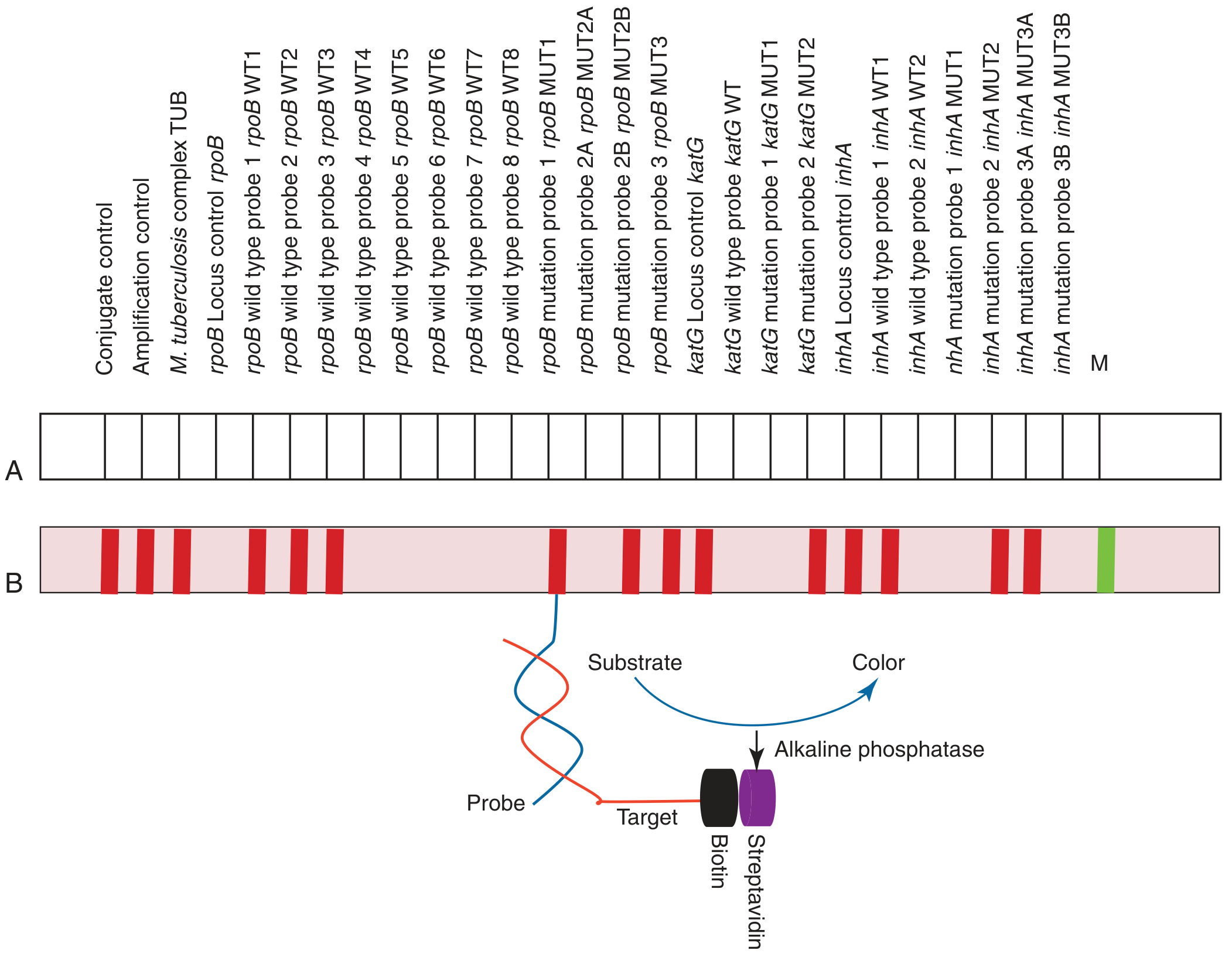
Configuration of GenoType MTBDRplus V2 (Hain Lifescience GmbH): (A) probe map layout, (B) actual band pattern. Amplified targets hybridize to membrane-fixed probes; the biotin-streptavidin-alkaline phosphatase system generates a colorimetric signal. — Tietz Textbook of Laboratory Medicine, 7th Ed.
Components on the Strip (MTBDRplus as example)
| Band Type | Interpretation |
|---|---|
| Conjugate control | Confirms streptavidin conjugate is functional |
| Amplification control (AC) | Confirms PCR amplification was successful |
| M. tuberculosis complex (TUB) | Species identification |
| Wild-type (WT) probes | Detect the normal (susceptible) sequence; presence = susceptible |
| Mutation (MUT) probes | Detect known resistance mutations; presence = resistant |
| Locus control | Internal control for each target gene locus |
Commercially Available LPAs for TB
| Assay | Manufacturer | Targets |
|---|---|---|
| GenoType MTBDRplus V2 | Hain Lifescience, Germany | rpoB (RIF resistance), katG + inhA (INH resistance) → detects MDR-TB |
| GenoType MTBDRsl | Hain Lifescience | gyrA/gyrB (fluoroquinolone), rrs + eis (aminoglycosides) → detects XDR-TB |
| INNO-LiPA Rif.TB | Fujirebio (Innogenetics) | rpoB only |
| INNO-LiPA HCV | Fujirebio | Hepatitis C genotyping |
| INNO-LiPA HPV | Fujirebio | HPV genotyping |
Genetic Targets and Resistance Mutations Detected
(Harrison's Principles of Internal Medicine, 22nd Ed.)
| Drug | Gene | Mutation Detection Rate |
|---|---|---|
| Rifampicin (RIF) | rpoB (RNA polymerase β-subunit) | ~95% of resistant strains |
| Isoniazid (INH) | katG (catalase-peroxidase) | 50–95% |
| inhA promoter | up to 45% | |
| Pyrazinamide | pncA | up to 98% |
| Ethambutol | embB | 50–65% |
| Fluoroquinolones | gyrA/gyrB | 75–95% |
| Aminoglycosides | rrs, eis promoter | up to 80% |
Interpretation Logic:
- WT band present + MUT band absent → Susceptible
- WT band absent + MUT band present → Resistant (specific known mutation)
- WT band absent + MUT band absent → Resistant (unknown/novel mutation — requires sequencing)
- WT band present + MUT band present → Mixed infection or heteroresistance
Specimen Requirements
- Direct specimen: AFB smear-positive sputum (without prior culture)
- Indirect specimen: positive liquid or solid culture
- Processing requires Biosafety Level 2 (or Class II laminar flow cabinet minimum)
- DNA must be extracted with adequate decontamination
Performance Characteristics
| Parameter | Value |
|---|---|
| Sensitivity for RIF resistance | ≥95% (smear-positive specimens) |
| Specificity | ≥95% for both RIF and INH resistance |
| Sensitivity for INH resistance | Slightly lower (some mutations not covered) |
| Turnaround time | 1–2 days (vs. 5–7 days for liquid culture DST; vs. 28 days for agar proportion) |
Note: Specificity (drug susceptible by both LPA and phenotypic DST) is ≥95%; sensitivity (drug resistant by both) is somewhat lower. — Henry's Clinical Diagnosis and Management by Laboratory Methods
WHO Recommendations
The WHO endorses LPA as a rapid molecular test for:
- Screening of patients at risk of MDR-TB (prior treatment, contact of MDR case, high-burden settings)
- Use directly on AFB smear-positive sputum — WHO Policy 2008
- MTBDRsl for second-line drug resistance (pre-XDR and XDR-TB screening)
The MTBDRplus is not FDA-approved but is recommended by WHO and widely used in countries like South Africa. — Fishman's Pulmonary Diseases and Disorders, 2-Volume Set
Advantages
- Rapid — results in 24–48 hours from smear-positive specimen, far faster than culture-based DST (weeks)
- Simultaneous — detects organism identity AND multiple drug resistance mutations in one test
- Direct specimen use — can be applied to smear-positive sputum without prior culture
- High specificity — stringent hybridization conditions prevent cross-reactivity
- Multiplexing — many probes on a single strip
- Standardized — visual band pattern is semi-quantitative and interpretable with a reading card
- Cost-effective for high-burden settings compared to sequencing
Limitations
- Does not detect all resistance mutations — only covers known mutations; novel mutations (WT absent, MUT absent) require DNA sequencing
- Lower sensitivity in smear-negative or paucibacillary specimens — sensitivity drops to 40–60% in AFB smear-negative sputum
- Cannot assess phenotypic resistance level (e.g., low-level vs. high-level INH resistance)
- Does not replace culture — culture remains gold standard; LPA cannot determine viability or assess all drugs
- Requires PCR facility — biosafety considerations; cannot be used as a point-of-care test
- Resistance to pyrazinamide not detected by MTBDRplus (requires a separate assay)
- Mixed infection/contamination can give ambiguous results
LPA vs. Other Rapid Diagnostics
| Feature | LPA (MTBDRplus) | Xpert MTB/RIF Ultra | Phenotypic DST |
|---|---|---|---|
| Turnaround | 1–2 days | ~2 hours | 5–28 days |
| Drugs detected | RIF + INH (± second-line) | RIF only | Multiple drugs |
| Direct specimen | Smear+ only | Yes (smear+ and −) | Smear+ or culture |
| Point-of-care | No | Near (cartridge-based) | No |
| Novel mutations | Missed | Missed | Detected |
| WHO-endorsed | Yes | Yes | Yes (gold standard) |
Applications Beyond TB
- Hepatitis C virus (HCV) genotyping — INNO-LiPA HCV assay (genotypes 1–6)
- HIV drug resistance — detection of reverse transcriptase/protease mutations
- Hepatitis B — lamivudine resistance mutations
- HPV genotyping — high-risk vs. low-risk types
- Hepatitis C genotyping for treatment selection (e.g., direct-acting antivirals are genotype-dependent)
Summary for Exam
LPA = Reverse hybridization + PCR amplification + membrane-bound probes + colorimetric detectionKey assay: GenoType MTBDRplus — detects rpoB (RIF resistance) + katG/inhA (INH resistance) = MDR-TB diagnosis within 1–2 days from smear-positive sputumWild-type band loss = resistance; Mutation band appearance = specific known mutation; WHO-endorsed for MDR-TB screening
Sources:
- Tietz Textbook of Laboratory Medicine, 7th Edition (Hybridization Arrays, p. 2665–66)
- Henry's Clinical Diagnosis and Management by Laboratory Methods (Susceptibility Testing, p. 1366)
- Fishman's Pulmonary Diseases and Disorders, 2-Volume Set (Nucleic Acid Amplification, p. 2296)
- Harrison's Principles of Internal Medicine, 22nd Ed. (Drug-Resistant TB, p. 1441)
This is a shared conversation. Sign in to Orris to start your own chat.