Explain second and third answer for pharm d student 1st year
Reading File
Finding Sources
Finding Sources
Finding Sources
Reading File
Reading File
Reading File
Reading File
Finding Sources
Finding Sources
Loading Image
Loading Image
Finding Sources
Reading File
I now have comprehensive information from the textbooks. Let me compile the full answers for both questions.
Question 2: Factors Affecting Enzyme Activity + Properties of Enzymes
Properties of Enzymes
Before diving into factors, here are the key properties of enzymes you need to know:
| Property | Explanation |
|---|---|
| Proteins | All enzymes are proteins (except ribozymes — RNA-based). |
| Catalysts | Speed up reactions without being consumed or permanently changed. |
| Specificity | Each enzyme acts on only one or a few specific substrates (binding specificity). |
| Reaction specificity | Catalyze only one type of reaction — no by-products. |
| Stereospecificity | Recognize only one enantiomeric (mirror image) form of a substrate (e.g., proteases act only on L-amino acids). |
| Do NOT shift equilibrium | They speed up reaching equilibrium but don't change the equilibrium constant (Keq). |
| Regulated | Activity is controlled by allosteric molecules, inhibitors, or covalent modification. |
— Henry's Clinical Diagnosis and Management by Laboratory Methods
Factors Affecting Enzyme Activity
1. Temperature
- Every 10°C rise in temperature approximately doubles enzyme activity (Q10 effect).
- However, high temperatures cause denaturation — enzymes lose their 3D shape and activity.
- Example: Creatine kinase (CK) denatures at 37°C; amylase at 45°C.
- Exception: Taq polymerase remains stable at 95°C (used in PCR).
- Low temperatures slow reactions but preserve enzyme stability — useful for storing samples.
2. pH
- Each enzyme has an optimal pH at which it works best.
- Pepsin (stomach): optimal pH ~2 (acidic)
- Trypsin (small intestine): optimal pH ~8 (slightly alkaline)
- Most body enzymes: optimal pH ~7.4
- pH affects the ionization of amino acids in the active site, changing enzyme shape and substrate binding. Extreme pH causes denaturation.
3. Substrate Concentration ([S])
- As [S] increases, reaction velocity (v) increases — more enzyme-substrate complexes form.
- At very high [S], the enzyme becomes saturated and velocity reaches a maximum (V
max). - This relationship follows Michaelis-Menten kinetics: the Km (Michaelis constant) is the [S] at which velocity = ½ V
max. A lower Km means higher affinity for the substrate.
4. Enzyme Concentration
- If substrate is in excess, reaction rate is directly proportional to enzyme concentration — more enzyme molecules = more active sites = faster reaction.
5. Cofactors and Coenzymes
- Many enzymes require non-protein helpers:
- Cofactors: inorganic ions (Mg²⁺, Zn²⁺, Fe²⁺)
- Coenzymes: organic molecules (NAD⁺, FAD, Coenzyme A — most derived from vitamins)
6. Inhibitors (see Question 3 below)
Question 3: Enzyme Inhibition — Definition and Types
Definition
Any substance that decreases the velocity of an enzyme-catalyzed reaction is called an inhibitor.
Types of Enzyme Inhibition
A. Irreversible Inhibition
- Inhibitor binds to the enzyme via a covalent bond — permanent, cannot be reversed by dilution.
- Destroys enzyme function permanently.
- Example: Lead (Pb²⁺) irreversibly inhibits ferrochelatase (an enzyme in heme synthesis) by forming covalent bonds with cysteine's –SH group.
- Aspirin irreversibly inhibits cyclooxygenase (COX), blocking prostaglandin synthesis.
B. Reversible Inhibition
Inhibitor binds via non-covalent bonds — can be removed by dilution. Two main types:
1. Competitive Inhibition
- The inhibitor looks like the substrate and competes for the same active site.
- Substrate and inhibitor cannot bind simultaneously.
| Parameter | Effect |
|---|---|
| V | Unchanged — can be overcome by adding excess substrate |
| Km | Increased (apparent) — more substrate needed to reach ½ V |
Lineweaver-Burk plot: Lines intersect on the y-axis (same 1/Vmax, different x-intercepts).
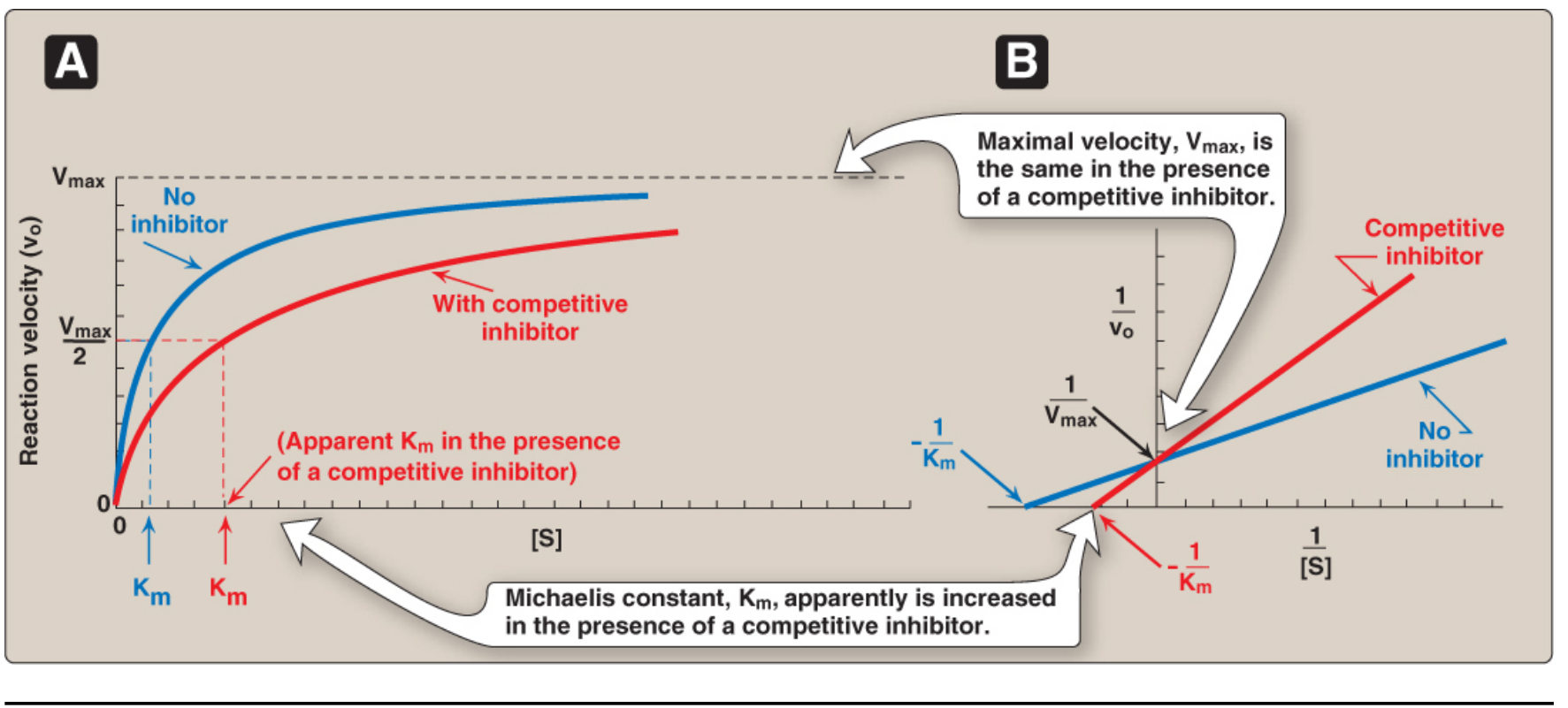
Pharma example: Statins (e.g., pravastatin, atorvastatin) competitively inhibit HMG-CoA reductase, blocking cholesterol synthesis. Methotrexate competitively inhibits dihydrofolate reductase (DHFR).
2. Noncompetitive Inhibition
- The inhibitor binds to a different site (allosteric site) — not the active site.
- Can bind to free enzyme OR the enzyme-substrate (ES) complex.
- Substrate binding is unaffected, but the reaction cannot proceed normally.
| Parameter | Effect |
|---|---|
| V | Decreased — cannot be overcome by adding more substrate |
| Km | Unchanged — substrate affinity is unaffected |
Lineweaver-Burk plot: Lines intersect on the x-axis (same –1/Km, different y-intercepts).
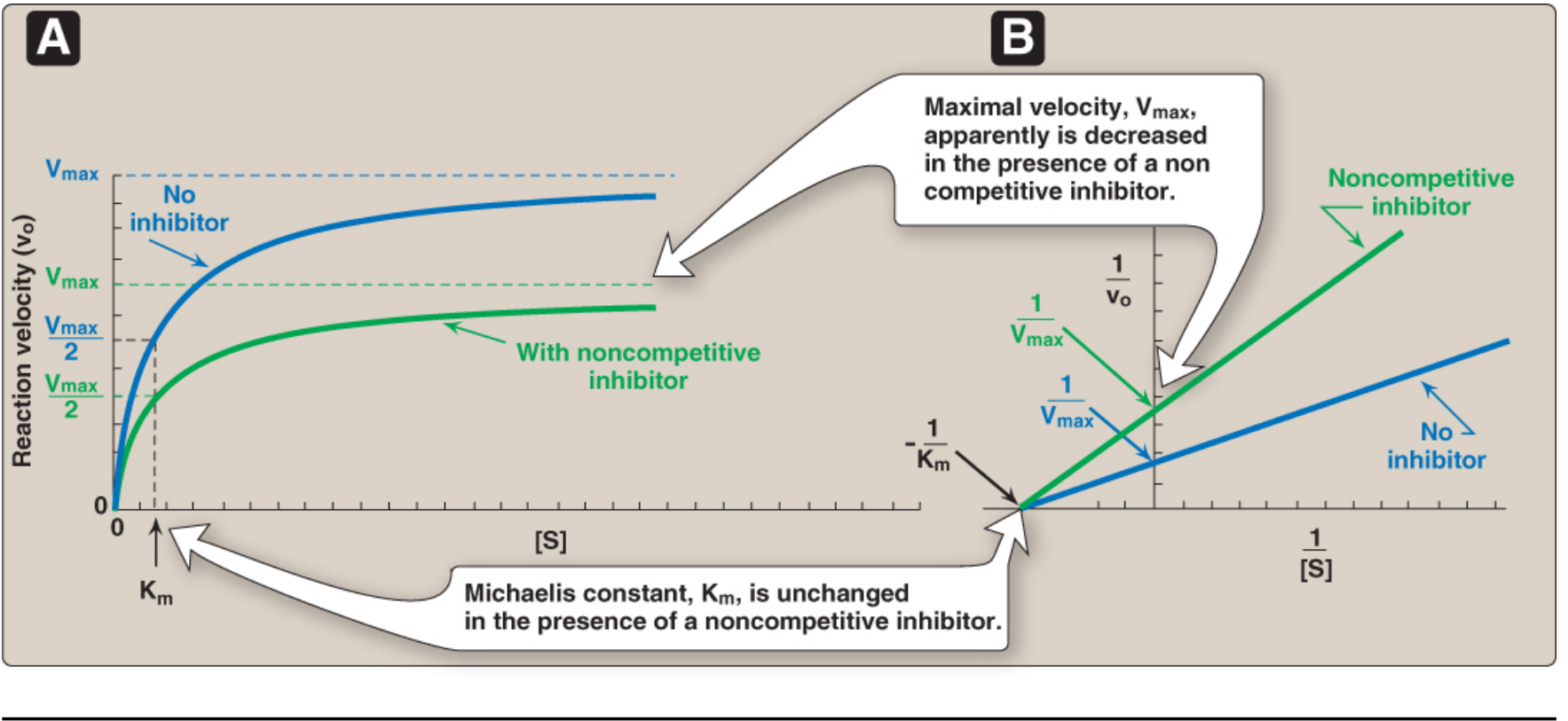
Example: Heavy metals like Hg²⁺ acting on –SH groups away from the active site.
3. Uncompetitive Inhibition
- Inhibitor binds only to the ES complex (not free enzyme).
- Both V
maxand Km are decreased proportionally. - Lineweaver-Burk plot: Parallel lines (same slope, different intercepts).
Quick Comparison Table
| Feature | Competitive | Noncompetitive | Uncompetitive |
|---|---|---|---|
| Binding site | Active site | Allosteric site | ES complex only |
| V | Unchanged | Decreased | Decreased |
| Km | Increased | Unchanged | Decreased |
| Reversible by excess [S]? | Yes | No | No |
| Lineweaver-Burk | Same y-intercept | Same x-intercept | Parallel lines |
Drug Examples (Important for Pharm D)
| Drug | Mechanism |
|---|---|
| Statins (atorvastatin) | Competitive inhibition of HMG-CoA reductase |
| Penicillin / Amoxicillin | Irreversible inhibition of bacterial cell wall synthesis enzymes |
| ACE inhibitors (captopril, enalapril) | Inhibit angiotensin-converting enzyme → vasodilation |
| Aspirin | Irreversible inhibition of COX (cyclooxygenase) |
| Methotrexate | Competitive inhibition of DHFR |
Sources: Lippincott's Illustrated Reviews Biochemistry 8th ed.; Henry's Clinical Diagnosis and Management by Laboratory Methods
Explain the factor affecting enzyme activity. Add a note on the properites of enzymes for pharm d 1st year
Reading File
Finding Sources
Finding Sources
Reading File
Reading File
Reading File
Reading File
Finding Sources
Finding Sources
Finding Sources
Reading File
Searching Images
factors affecting enzyme activity pH temperature substrate concentration graph
Loading Image
Finding Sources
I now have comprehensive content from the textbooks. Here is the full, exam-ready answer:
Factors Affecting Enzyme Activity + Properties of Enzymes
(Pharm D 1st Year — Biochemistry)
PART 1: FACTORS AFFECTING ENZYME ACTIVITY
Enzyme activity refers to the rate at which an enzyme catalyzes its reaction. Several factors can increase or decrease this rate:
1. 🌡️ Temperature
- Enzyme activity increases as temperature rises — generally, every 10°C increase doubles the reaction rate (Q₁₀ effect).
- This is because higher temperature increases molecular kinetic energy, leading to more frequent enzyme-substrate collisions.
- However, above an optimal temperature, the enzyme begins to denature — its 3D structure unfolds, active site is destroyed, and activity drops sharply to zero.
Key examples:
| Enzyme | Starts to Denature At |
|---|---|
| Creatine Kinase (CK) | 37°C |
| Amylase | 45°C |
| Taq Polymerase | Stable up to 95°C (used in PCR) |
Practical note: Clinical lab samples stored at 4°C remain stable for ~1 week. Deeper freezing (−80°C) preserves enzyme activity longer.
2. 🧪 pH
- Every enzyme has an optimum pH at which it shows maximum activity.
- Changes in pH affect the ionization of amino acid residues in the active site, altering the enzyme's shape and ability to bind substrate.
- Extreme pH values on either side cause denaturation.
Key examples:
| Enzyme | Optimal pH | Location |
|---|---|---|
| Pepsin | ~2 (acidic) | Stomach |
| Trypsin | ~8 (basic) | Small intestine |
| Alkaline Phosphatase (ALP) | 9–10 | Liver/bone |
| Most body enzymes | ~7.4 | Blood/cytoplasm |
3. 📊 Substrate Concentration [S]
- As substrate concentration increases, reaction velocity (v) increases — more substrates fill available active sites.
- At very high [S], all active sites are occupied → the enzyme is saturated → velocity reaches a maximum (V
max). - This follows Michaelis-Menten kinetics:
$$v_0 = \frac{V_{max}[S]}{K_m + [S]}$$
- K
m(Michaelis constant) = the [S] at which v = ½ Vmax- Low K
m= high affinity for substrate (less substrate needed) - High K
m= low affinity for substrate
- Low K
4. ⚗️ Enzyme Concentration [E]
- When substrate is in excess, reaction rate is directly proportional to enzyme concentration.
- More enzyme molecules = more active sites available = faster product formation.
- Clinically, enzyme concentration in blood is measured to diagnose disease (e.g., elevated ALT/AST in liver damage).
5. 🔩 Cofactors and Coenzymes (Activators)
Many enzymes are inactive alone and require non-protein helpers:
| Type | Description | Examples |
|---|---|---|
| Cofactors | Inorganic metal ions | Zn²⁺, Mg²⁺, Fe²⁺, Cu²⁺ |
| Coenzymes | Small organic molecules (loosely bound) | NAD⁺ (from niacin/Vit B₃), FAD (from riboflavin/Vit B₂) |
| Prosthetic groups | Coenzymes permanently bound to enzyme | FAD, heme |
- The complete, active enzyme (protein + cofactor/coenzyme) = Holoenzyme
- The inactive protein part alone = Apoenzyme
- Most coenzymes are derived from B-vitamins — this is why B-vitamin deficiency causes metabolic disease.
Holoenzyme = Apoenzyme + Cofactor/Coenzyme
6. 🚫 Inhibitors
Any substance that decreases enzyme activity is an inhibitor. (Covered in depth in Question 3.) Key types:
- Irreversible — covalent binding (e.g., aspirin, lead)
- Reversible — competitive, noncompetitive, uncompetitive
7. 🧂 Ionic Strength / Salt Concentration
- High ionic strength decreases enzyme activity by disrupting the ionic interactions that stabilize enzyme structure.
- Dilution of clinical samples with appropriate enzyme diluents (containing plasma proteins like albumin) is necessary to maintain accurate enzyme measurements.
8. ⚡ Allosteric Regulation
- Small molecules called allosteric effectors bind to a site other than the active site.
- They cause conformational changes in the enzyme:
- Allosteric activators → increase activity
- Allosteric inhibitors → decrease activity
- This is the main mechanism for feedback regulation in metabolic pathways.
PART 2: NOTE ON PROPERTIES OF ENZYMES
What Are Enzymes?
Enzymes are biological catalysts — they speed up chemical reactions in living organisms without being permanently changed or consumed in the process.
Key Properties
1. Proteins in Nature
All enzymes are proteins (made of amino acids), EXCEPT ribozymes (RNA molecules with catalytic activity — e.g., ribosomes during peptide bond formation).
2. Catalytic Function
- Enzymes speed up reactions by lowering the activation energy (E
a) — the energy needed to start the reaction. - They do NOT change the equilibrium of a reaction (the final ratio of products to reactants is unchanged — only the speed of reaching equilibrium changes).
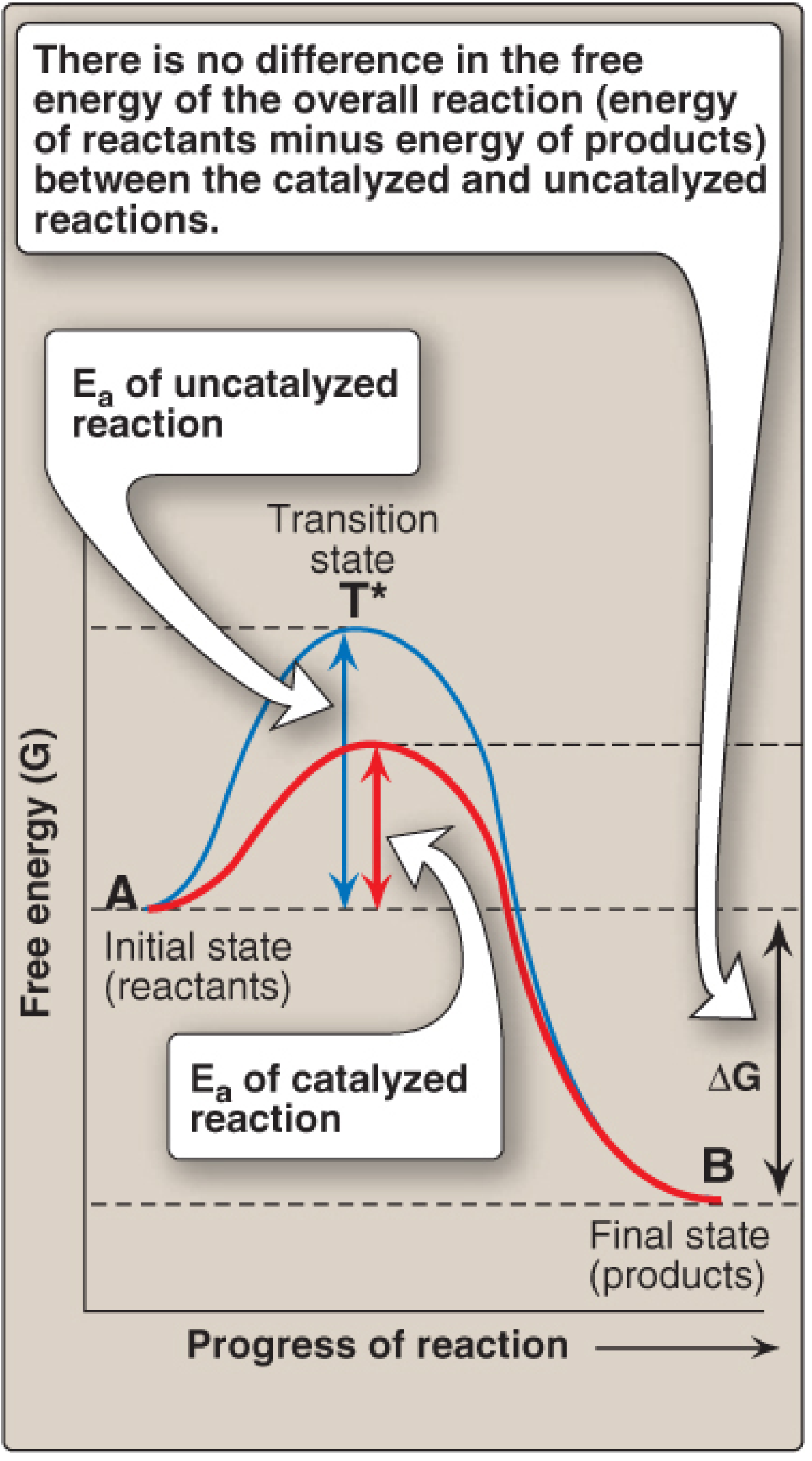
3. High Specificity
- Binding specificity: Each enzyme binds only one or a few specific substrates.
- Reaction specificity: Each enzyme catalyzes only one type of chemical reaction — virtually no by-products are formed.
- Stereospecificity: Enzymes recognize only one mirror-image (enantiomeric) form of a substrate. Example: Proteases act only on L-amino acids (not D-amino acids).
4. Active Site
- The active site is a small region on the enzyme where the substrate binds and the reaction occurs.
- Two models explain enzyme-substrate binding:
- Lock and Key model: Active site is a rigid, pre-formed shape that perfectly fits the substrate.
- Induced Fit model (more accepted): Active site is flexible; it changes shape when the substrate binds to achieve the best fit.
5. Not Consumed in Reaction
- Enzymes are regenerated after each reaction cycle — one enzyme molecule can process thousands of substrate molecules per second.
6. Regulated
- Enzyme activity can be turned up or down based on cellular needs via:
- Allosteric regulators
- Covalent modification (e.g., phosphorylation)
- Feedback inhibition (product inhibits early enzyme in a pathway)
- Induction/repression of gene expression
7. Cellular Compartmentalization
- Enzymes are localized in specific organelles. Example:
- Krebs cycle enzymes → mitochondria
- Digestive enzymes → lysosomes
- Glycolytic enzymes → cytoplasm
- This isolates reactions, prevents interference, and increases efficiency.
8. Cofactor Requirement
- Some enzymes need non-protein helpers (cofactors or coenzymes) for activity — without them, they are inactive (apoenzyme form).
9. Enzyme Specificity as Biomarkers
- Because many enzymes are tissue-specific, elevated levels in blood indicate organ damage.
- Elevated ALT/AST → liver disease
- Elevated CK-MB → myocardial infarction (heart attack)
- Elevated amylase/lipase → pancreatitis
Summary Table
| Property | Key Point |
|---|---|
| Chemical nature | Proteins (except ribozymes) |
| Function | Biological catalysts — lower activation energy |
| Consumed? | No — regenerated after each reaction |
| Change equilibrium? | No — only speed up reaching it |
| Specificity | High — one enzyme, one substrate, one reaction |
| Temperature effect | ↑ activity up to optimum → denaturation |
| pH effect | Optimum pH varies per enzyme |
| Cofactors | Required by many enzymes; coenzymes often from vitamins |
| Regulation | Allosteric, covalent modification, feedback inhibition |
Sources: Henry's Clinical Diagnosis and Management by Laboratory Methods; Lippincott's Illustrated Reviews: Biochemistry, 8th Ed.
This is a shared conversation. Sign in to Orris to start your own chat.