Give me an overview about protein separation technique for my biochemistry
Reading File
Finding Sources
Finding Sources
Reading File
Finding Sources
Reading File
Finding Sources
Reading File
Finding Sources
Finding Sources
Reading File
Reading File
Reading File
Loading Image
I now have comprehensive information from the textbooks. Let me compile the full overview.
Protein Separation Techniques — Biochemistry Overview
Protein separation exploits differences in charge, size, hydrophobicity, and molecular affinity. The major techniques fall into two broad categories: electrophoresis and column separations, with chemical precipitation as a simpler adjunct.
1. Electrophoresis
Electrophoresis moves proteins through a support medium under an electric field. The rate and direction of migration depend primarily on the net charge of each protein at the working pH (usually pH 8.6 in the clinical lab).
1a. Zone Electrophoresis (Agarose / Cellulose Acetate)
- Sample is applied to an agarose gel or cellulose acetate membrane.
- A voltage is applied for ~30 minutes; proteins migrate toward the anode.
- After migration, the gel is fixed (acetic acid), stained, then scanned by densitometry.
- Separates serum proteins into 5 classic fractions: albumin, α₁-, α₂-, β-, and γ-globulins.
- Electro-osmosis (endosmosis): when the support medium carries a negative charge, buffer flows toward the cathode, carrying proteins mechanically — this must be accounted for in interpretation.
Clinical use: Screening for monoclonal gammopathies, liver disease, inflammatory states, nephrotic syndrome.
Normal serum electrophoresis — proteins migrate from anode (top) to cathode:
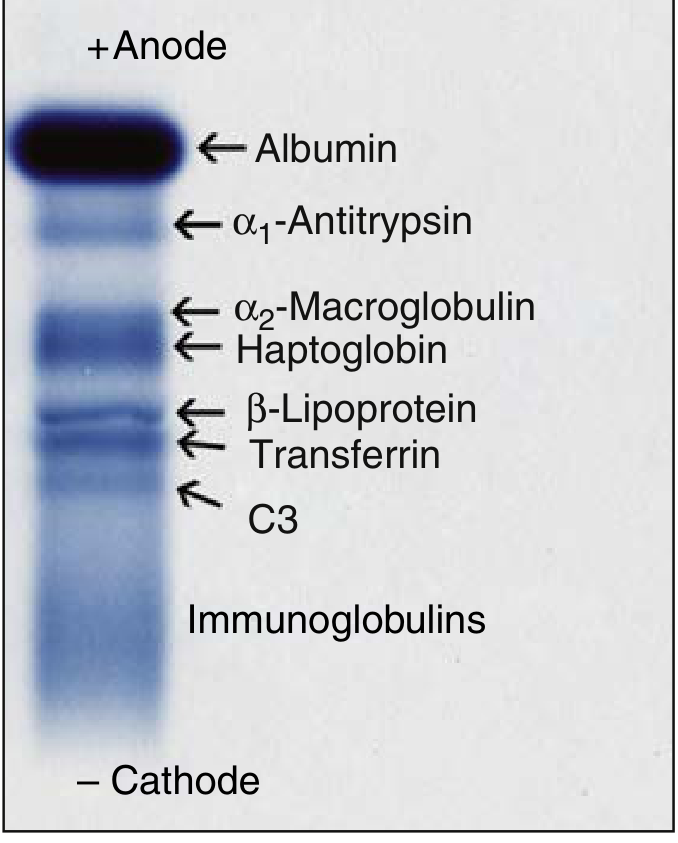
Figure 20.1 — Positions of major serum proteins in a normal person using electrophoresis (Henry's Clinical Diagnosis and Management by Laboratory Methods)
1b. SDS-PAGE (Sodium Dodecyl Sulfate–Polyacrylamide Gel Electrophoresis)
- Polyacrylamide is the support medium; porosity is adjustable by varying acrylamide concentration before polymerization.
- Proteins are denatured with SDS, which binds uniformly to the polypeptide chain and confers a large negative charge proportional to mass — abolishing native charge differences.
- Migration is controlled almost entirely by molecular weight (size): smaller proteins travel farther.
- Most widely used electrophoretic technique in molecular biology research.
- Not routinely used in the clinical lab due to its high resolution creating complex multitude of sub-bands.
1c. Isoelectric Focusing (IEF)
- Gel contains a pH gradient created by carrier ampholytes.
- Each protein migrates until it reaches the gel position where pH = its isoelectric point (pI) — at which the net charge is zero and migration stops.
- Separates proteins differing by very small charge differences (e.g., isoforms from posttranslational modifications like glycosylation or phosphorylation).
- Superior resolution for closely migrating proteins compared to standard zone electrophoresis.
1d. Two-Dimensional Gel Electrophoresis (2-DE)
- Combines IEF (1st dimension, separates by pI) and SDS-PAGE (2nd dimension, separates by molecular weight) at right angles.
- Can resolve hundreds of distinct protein spots in a complex mixture.
- Used in proteomics research; holds promise for diagnostic pattern analysis.
1e. Capillary Electrophoresis (CE)
- Separation occurs through a narrow capillary tube; similar in principle to agarose electrophoresis but automated.
- Detection at the effluent end quantitates protein without staining/scanning.
- Fast, highly quantitative, low reagent cost (though high equipment cost).
- For suspected monoclonal proteins: aliquots are pre-treated with specific antisera (anti-IgG, anti-IgA, anti-IgM, anti-κ, anti-λ) and re-run; the band that disappears identifies the monoclonal type.
2. Column Separations (Chromatography)
2a. Ion-Exchange Chromatography
Exploits ionic interactions between proteins and a charged stationary phase.
| Type | Column Matrix | Starting Condition | Elution |
|---|---|---|---|
| Anion-exchange | Positively charged (e.g., DEAE-cellulose) | Alkaline pH → proteins negatively charged | High salt or lower pH |
| Cation-exchange | Negatively charged (e.g., carboxymethylcellulose) | Acid pH → proteins positively charged (cations) | High salt or higher pH |
In cation-exchange, albumin elutes first, followed by α₁-, α₂-, β-, and γ-globulins.
Clinical example: Quantitation of glycosylated hemoglobin (HbA1c) using a dihydroxyboronate affinity matrix.
2b. Affinity Chromatography
- Based on specific, reversible binding between the protein of interest and a ligand covalently attached to the column matrix.
- Achieves a very high degree of purification in a single step.
- Bound protein is eluted by high salt, pH change, or denaturant (e.g., urea).
Examples:
- Anti-vWF antibody column → purifies factor VIII/vWF complex for transfusion therapy
- Protein A (from Staphylococcus aureus) column → purifies immunoglobulins
- Dye-coupled agarose → selectively binds albumin
- Dihydroxyboronate column → captures glycosylated hemoglobin (HbA1c measurement)
2c. Hydrophobic Interaction Chromatography (HIC)
- Column matrix interacts with hydrophobic regions of proteins.
- Sample applied at high salt (which enhances hydrophobic interactions); eluted with low salt.
- Complements ion-exchange chromatography well (since ion exchange elutes with high salt, directly usable as load for HIC).
2d. Size-Exclusion (Gel Filtration) Chromatography
- Separates proteins by molecular size through a porous bead matrix.
- Larger proteins cannot enter the pores → elute first.
- Smaller proteins enter the pores → elute later.
- Used for desalting, buffer exchange, and molecular weight estimation.
3. Chemical Precipitation
A simpler, older method:
- Adding ammonium sulfate, sodium sulfate, or methanol selectively precipitates globulins, leaving albumin in solution.
- Allows rough fractionation into albumin vs. globulin fractions.
- Total protein is then measured spectrophotometrically (biuret reagent, Coomassie blue) or by dye binding (bromocresol green/purple specifically binds albumin).
4. Immunological Methods (Complement to Separation)
After electrophoretic separation, immunological quantitation with specific antibodies identifies individual proteins:
- Nephelometry / turbidimetry — detects antibody-antigen precipitation; quantitates major serum proteins (transferrin, α₁-antitrypsin, IgG, etc.) on automated analyzers.
- Western Blotting (Immunoblotting) — proteins separated by SDS-PAGE are transferred to a membrane, then probed with specific antibodies; identifies molecular weight and abundance of a target protein.
- Immunoprecipitation / Co-immunoprecipitation — isolates a specific protein (and its binding partners) from complex lysates using antibody-conjugated beads, followed by SDS-PAGE or Western blot analysis.
Summary Comparison
| Technique | Basis of Separation | Resolution | Primary Use |
|---|---|---|---|
| Zone electrophoresis | Charge (pH 8.6) | 5 fractions | Clinical serum protein analysis |
| SDS-PAGE | Molecular weight | Very high | Research, MW determination |
| IEF | Isoelectric point | Very high | Protein isoforms, 2-DE |
| 2-DE | pI + MW | Highest | Proteomics |
| Capillary electrophoresis | Charge/size | High | Automated clinical labs |
| Ion-exchange chromatography | Ionic charge | Moderate–high | Protein fractionation, HbA1c |
| Affinity chromatography | Biospecific binding | Very high (single step) | Purification, therapeutics |
| Hydrophobic chromatography | Hydrophobicity | Moderate | Protein purification |
| Size-exclusion chromatography | Molecular size | Moderate | MW estimation, desalting |
| Chemical precipitation | Solubility | Low | Rough albumin/globulin split |
Sources: Henry's Clinical Diagnosis and Management by Laboratory Methods; Janeway's Immunobiology 10e; Cellular and Molecular Immunology
This is a shared conversation. Sign in to Orris to start your own chat.